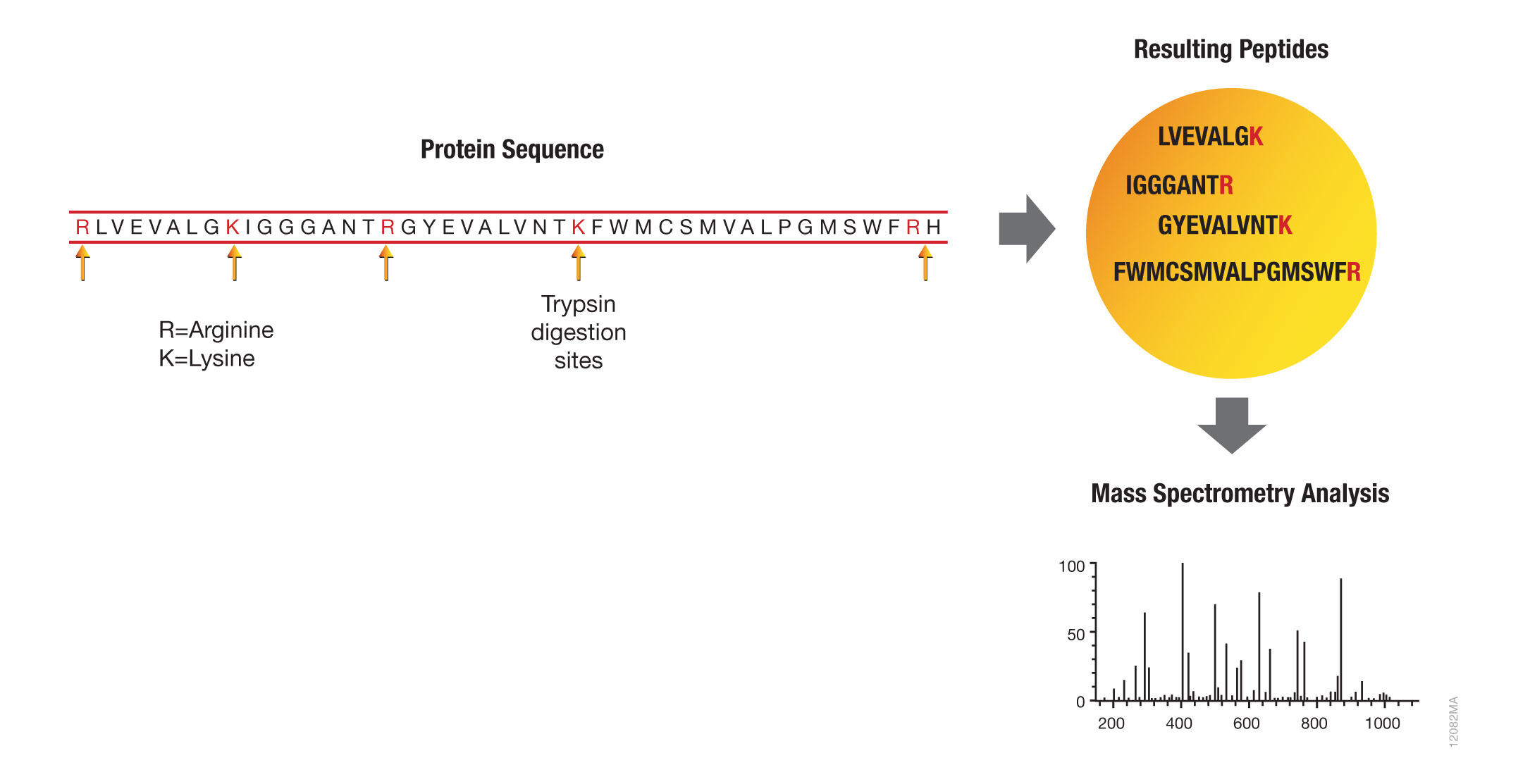
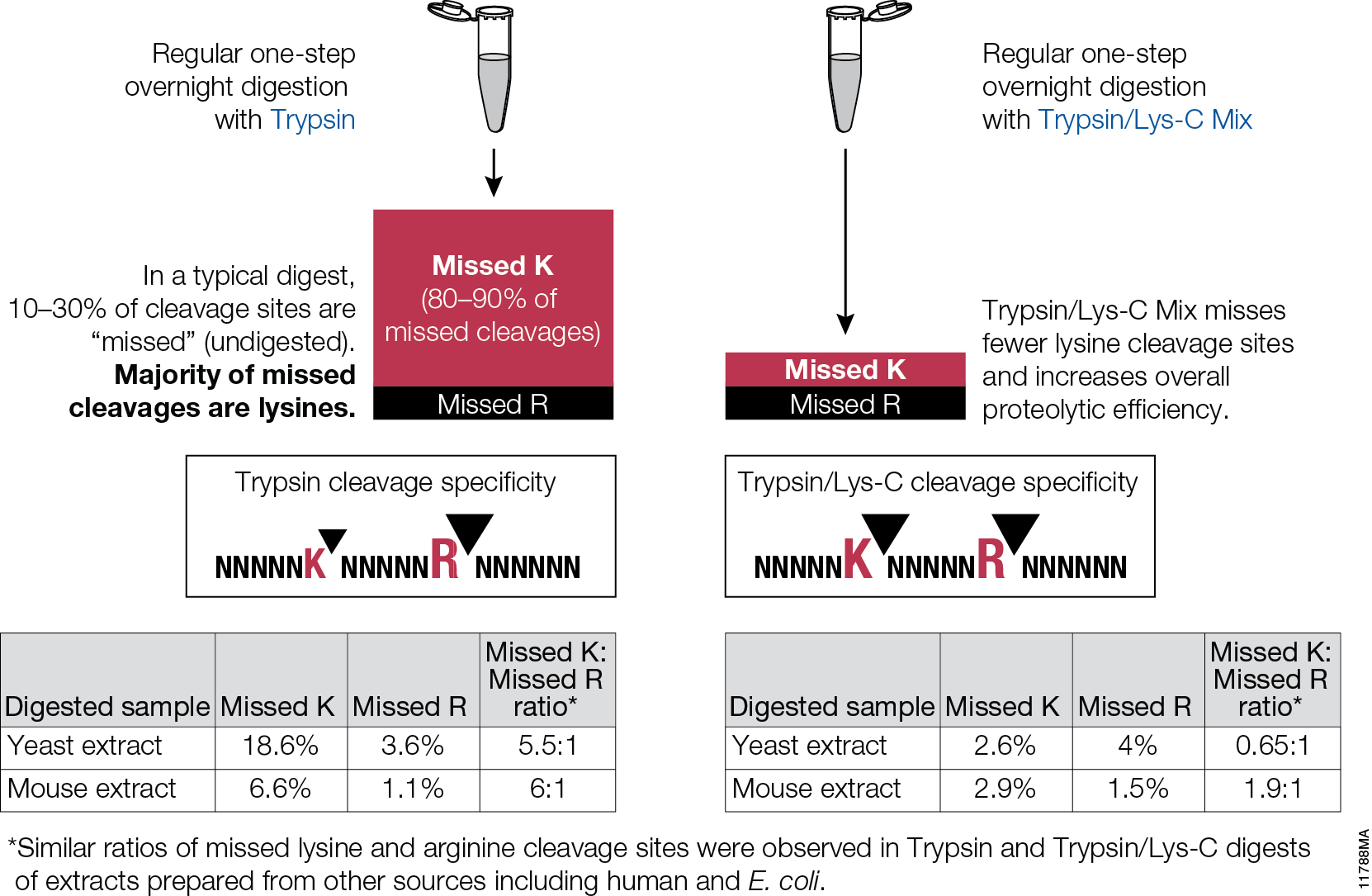
Prediction of Missed Proteolytic Cleavages for the Selection of Surrogate Peptides for Quantitative Proteomics | OMICS: A Journal of Integrative Biology
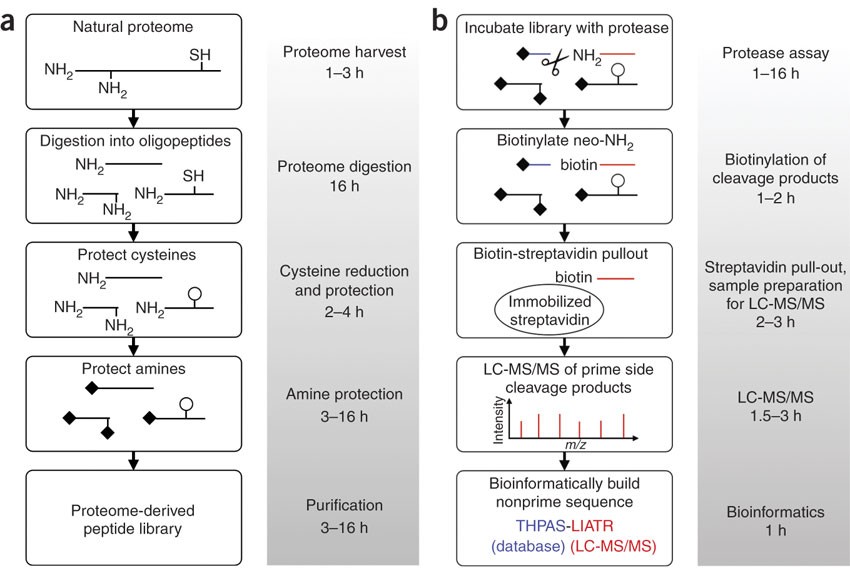
Proteome-derived, database-searchable peptide libraries for identifying protease cleavage sites | Nature Biotechnology

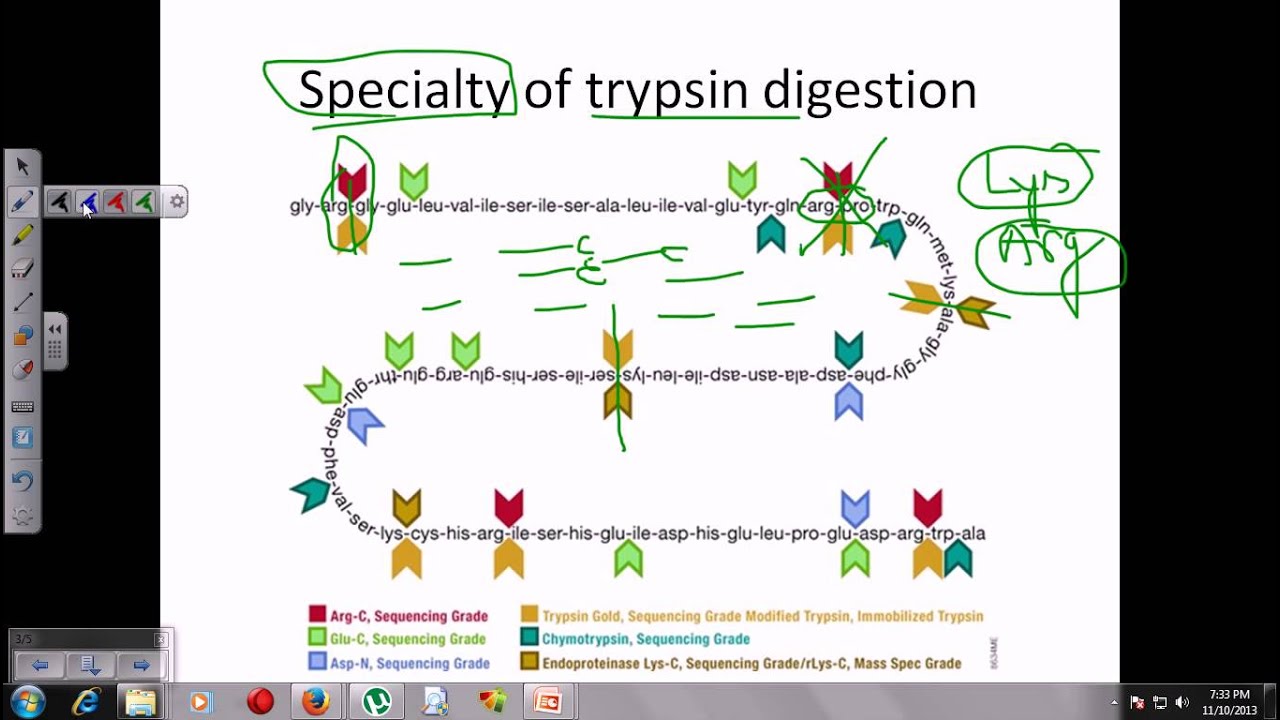
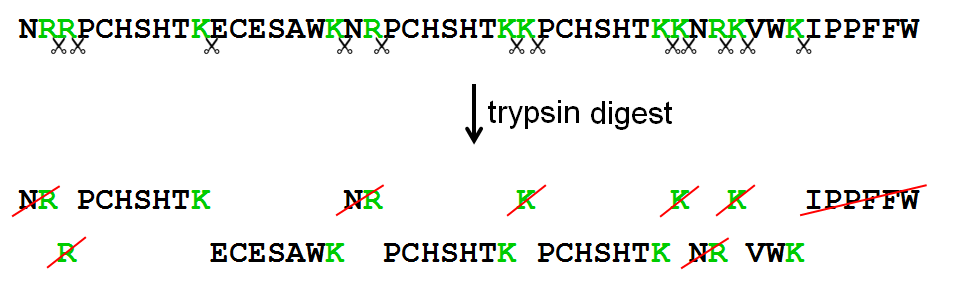
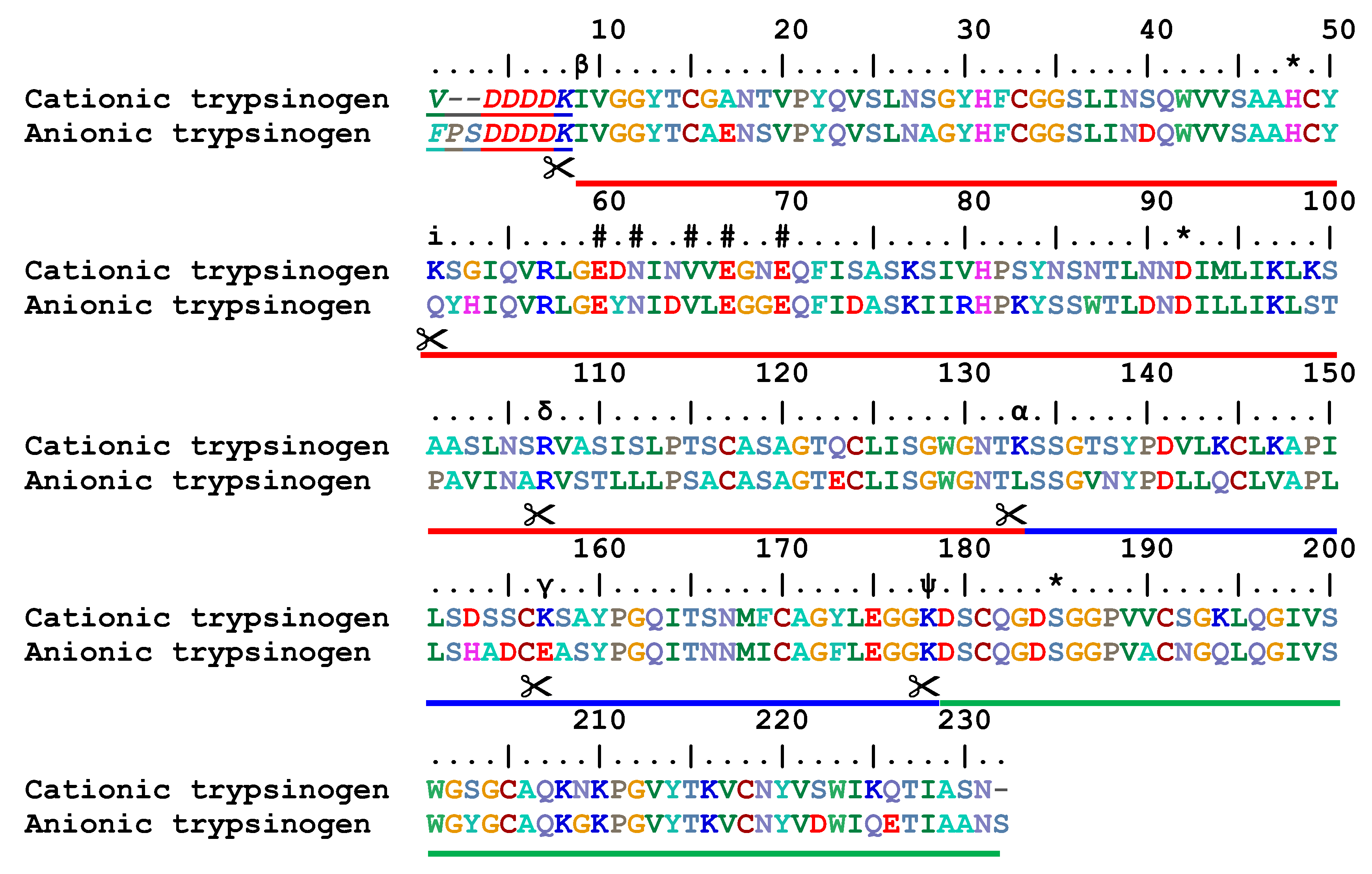
Schematic representation of the major trypsin cleavage sites and one... | Download Scientific Diagram

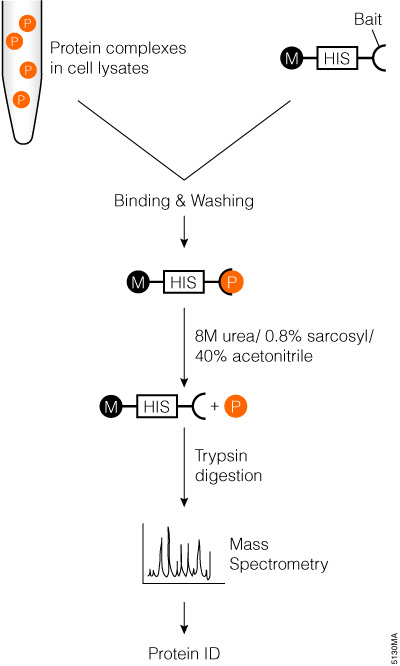
Proteomic Identification of protease Cleavage Sites (PICS) workflow.... | Download Scientific Diagram
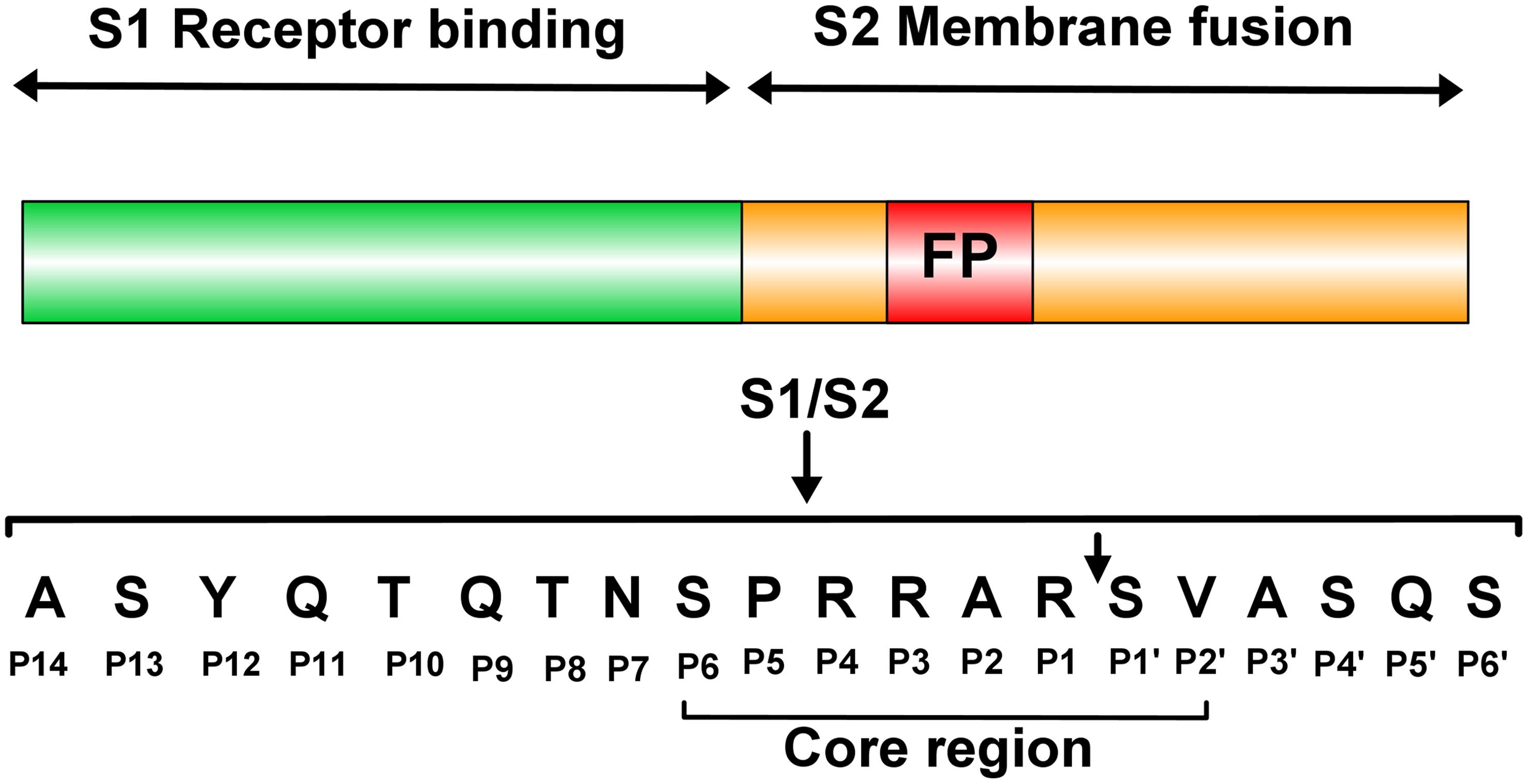
Hypertonic Saline and Aprotinin Inhibit Furin and Nasal Protease to Reduce SARS-CoV-2 Specific Furin Site Cleavage Activity

Insight into Trypsin Miscleavage: Comparison of Kinetic Constants of Problematic Peptide Sequences | Analytical Chemistry
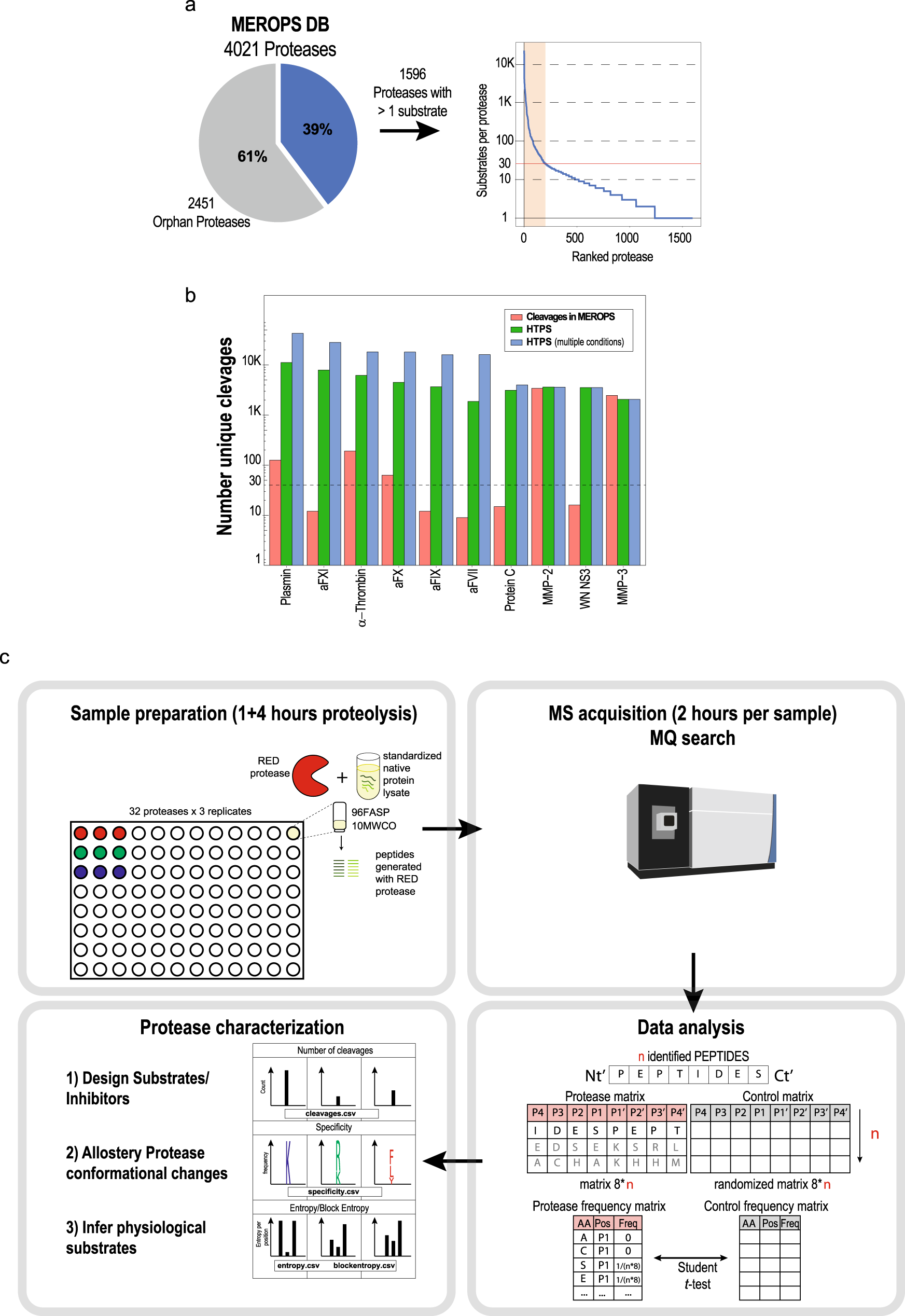
Mapping specificity, cleavage entropy, allosteric changes and substrates of blood proteases in a high-throughput screen | Nature Communications
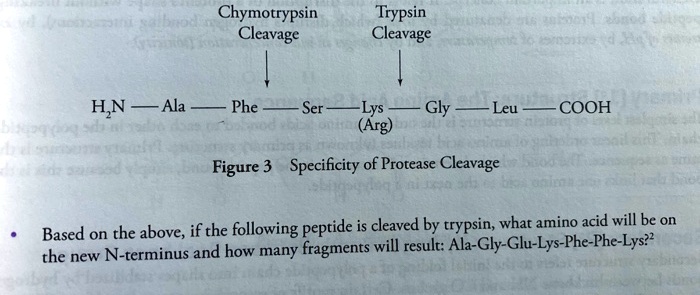
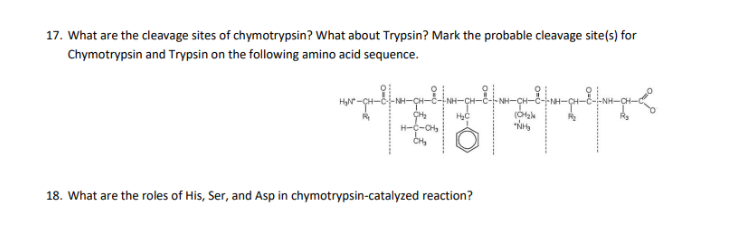
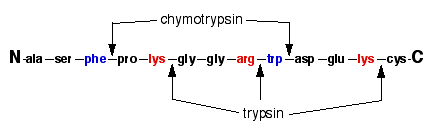
SOLVED: Chymotrypsin Cleavage Trypsin Cleavage HN Ala Phe Ser Lys (Arg) Gly Leu COOH Figure 3 Specificity of Protease Cleavage Based on the above; if the following peptide is cleaved by trypsin,

Global Sequencing of Proteolytic Cleavage Sites in Apoptosis by Specific Labeling of Protein N Termini: Cell

Protease cleavage site fingerprinting by label‐free in‐gel degradomics reveals pH‐dependent specificity switch of legumain | The EMBO Journal
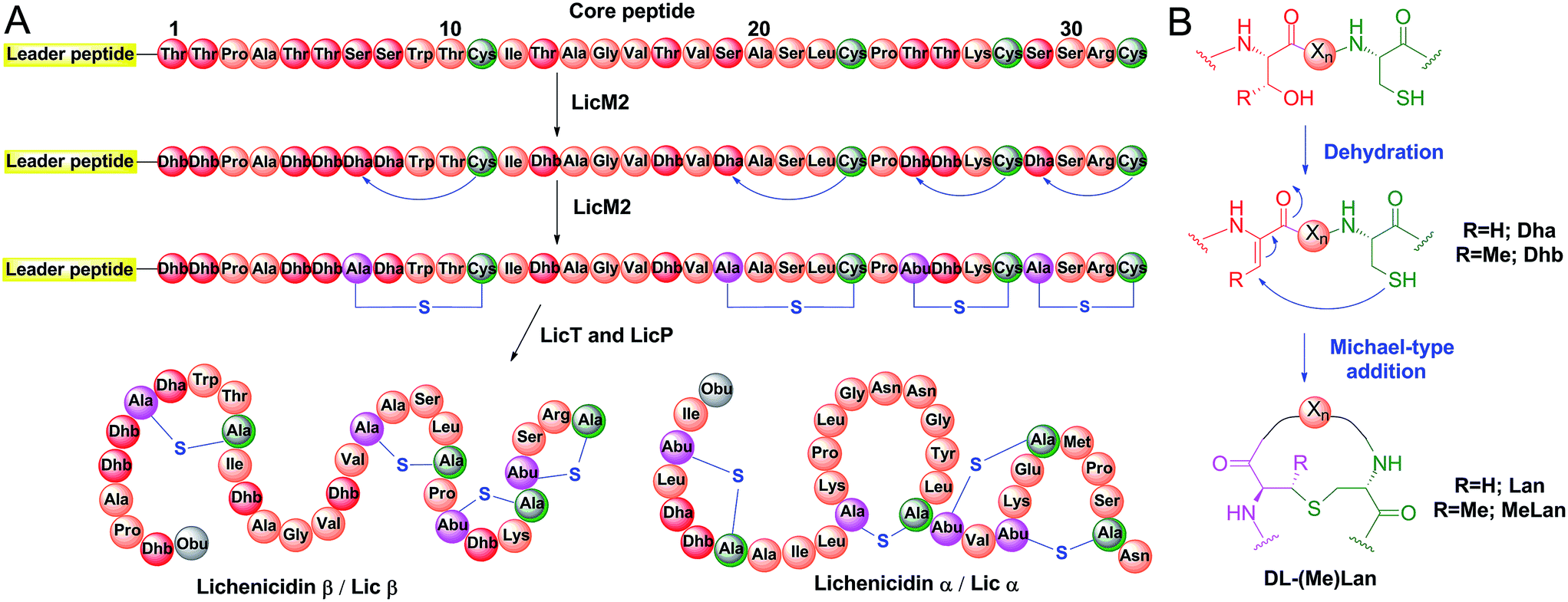
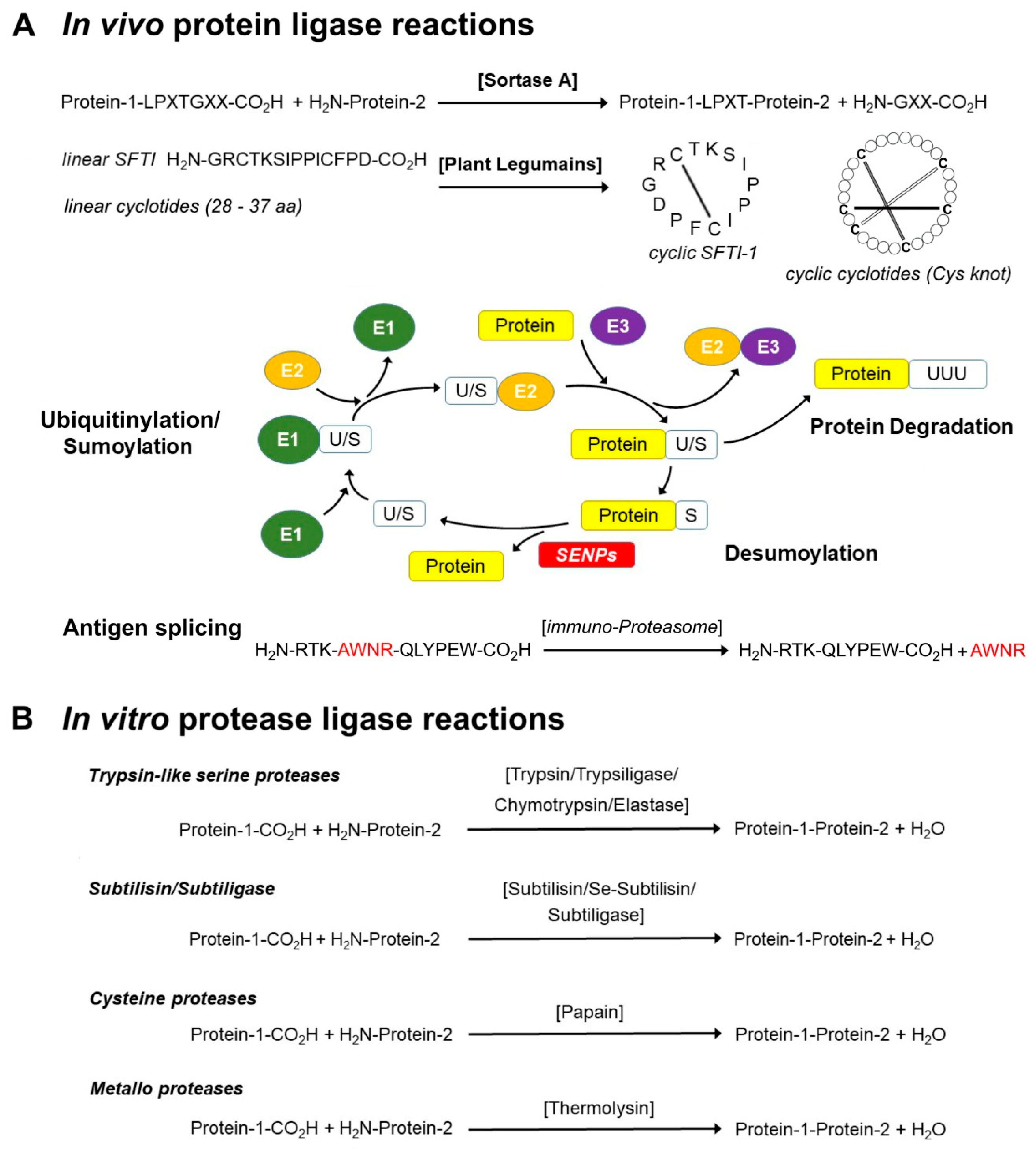
Applications of the class II lanthipeptide protease LicP for sequence-specific, traceless peptide bond cleavage - Chemical Science (RSC Publishing) DOI:10.1039/C5SC02329G