Phospho‐RNA‐seq: a modified small RNA‐seq method that reveals circulating mRNA and lncRNA fragments as potential biomarkers in human plasma | The EMBO Journal

Single-cell CAS-seq reveals a class of short PIWI-interacting RNAs in human oocytes | Nature Communications

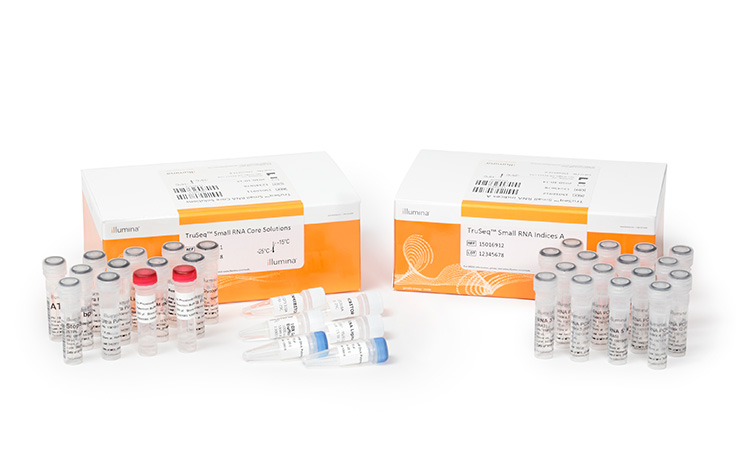
Incorporation of unique molecular identifiers (UMIs) in TruSeq adapters improves the accuracy of quantitative sequencing | RNA-Seq Blog

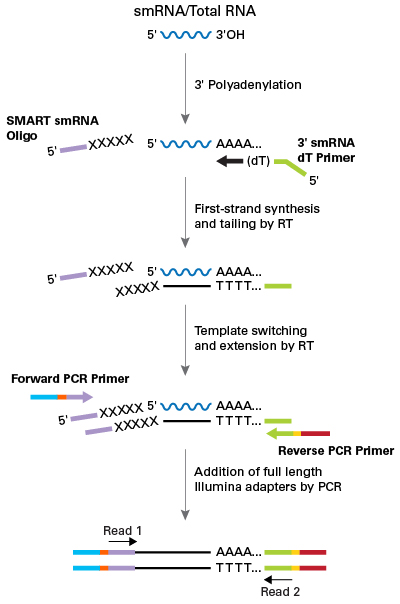
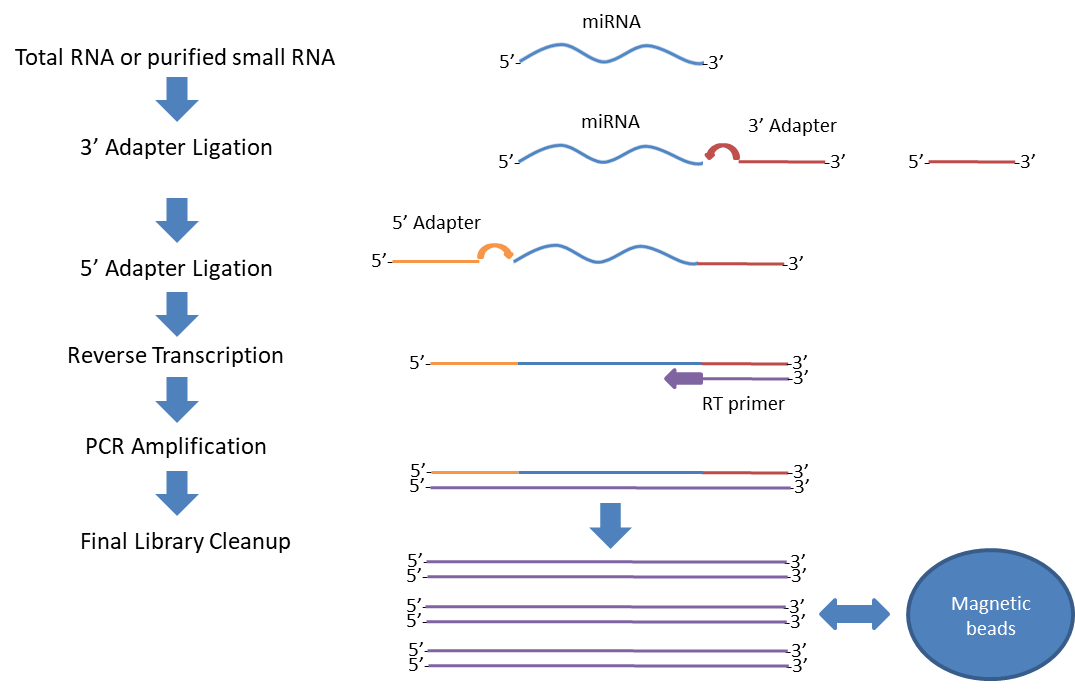
The protocol for preparing samples for small RNA sequencing. Total RNA... | Download Scientific Diagram
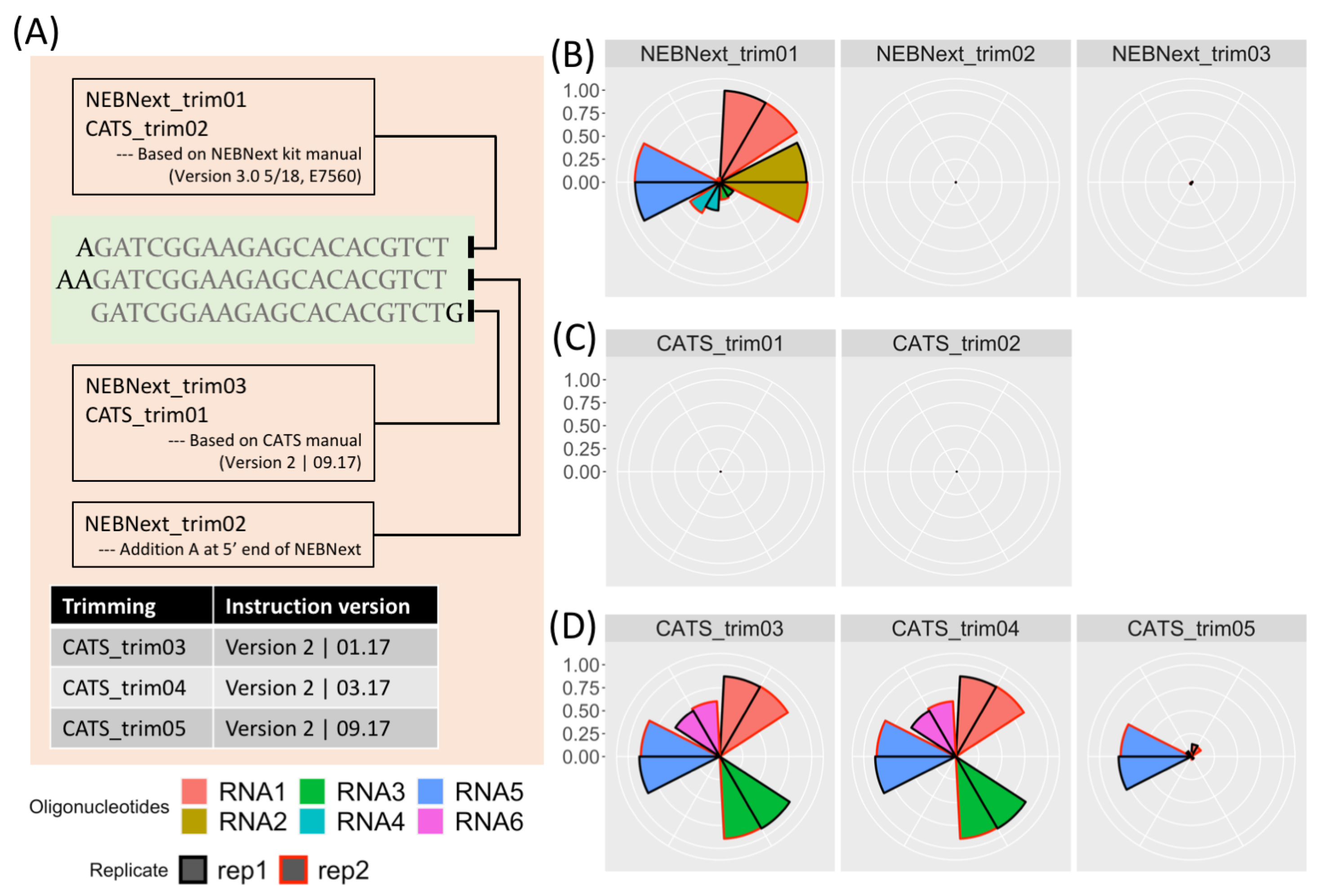
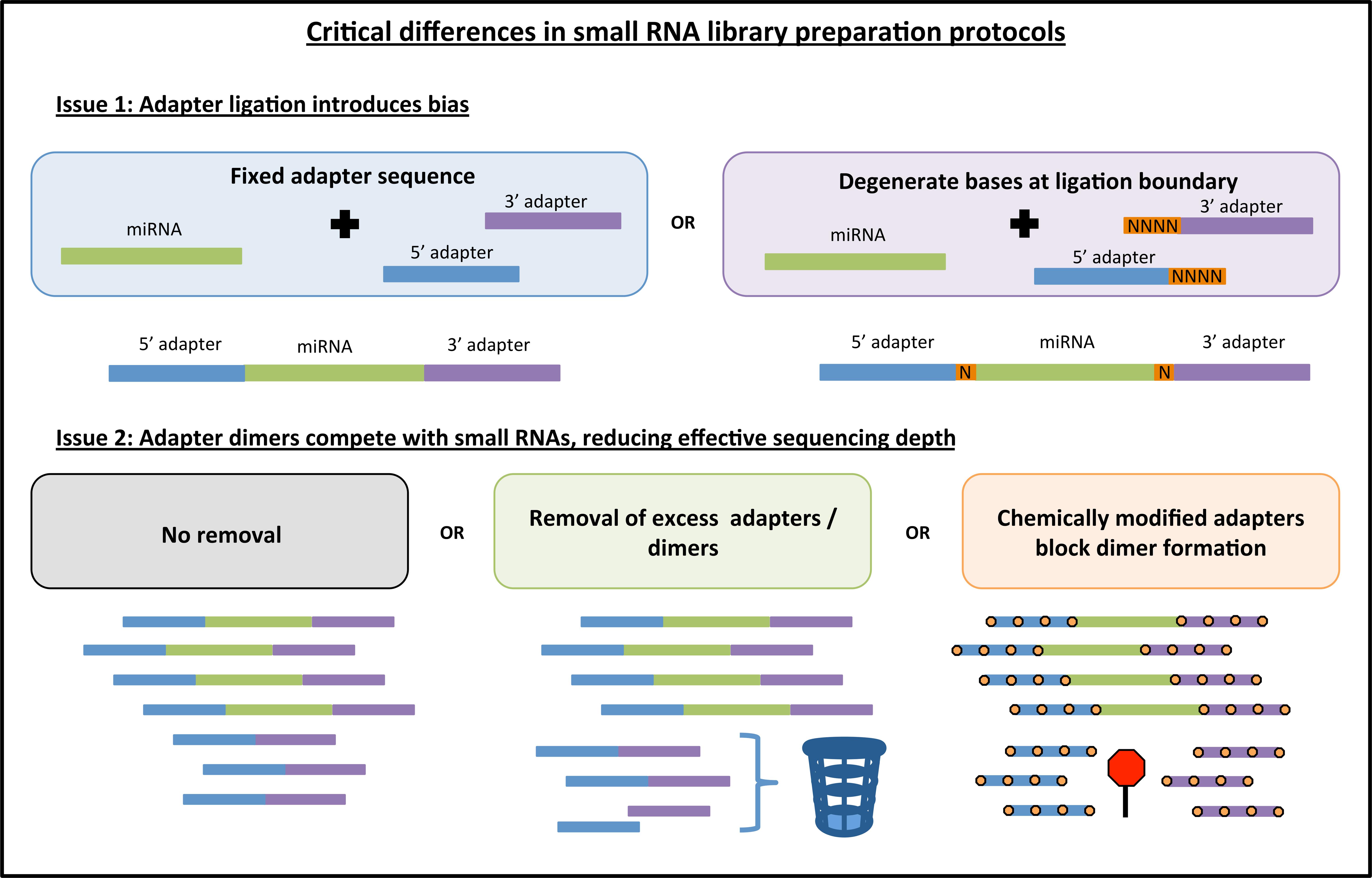
ncRNA | Free Full-Text | Accurate Adapter Information Is Crucial for Reproducibility and Reusability in Small RNA Seq Studies

Phospho‐RNA‐seq: a modified small RNA‐seq method that reveals circulating mRNA and lncRNA fragments as potential biomarkers in human plasma | The EMBO Journal
Quantitative Bias in Illumina TruSeq and a Novel Post Amplification Barcoding Strategy for Multiplexed DNA and Small RNA Deep Sequencing | PLOS ONE
Blocking oligo design and application workflows using modified TruSeq... | Download Scientific Diagram
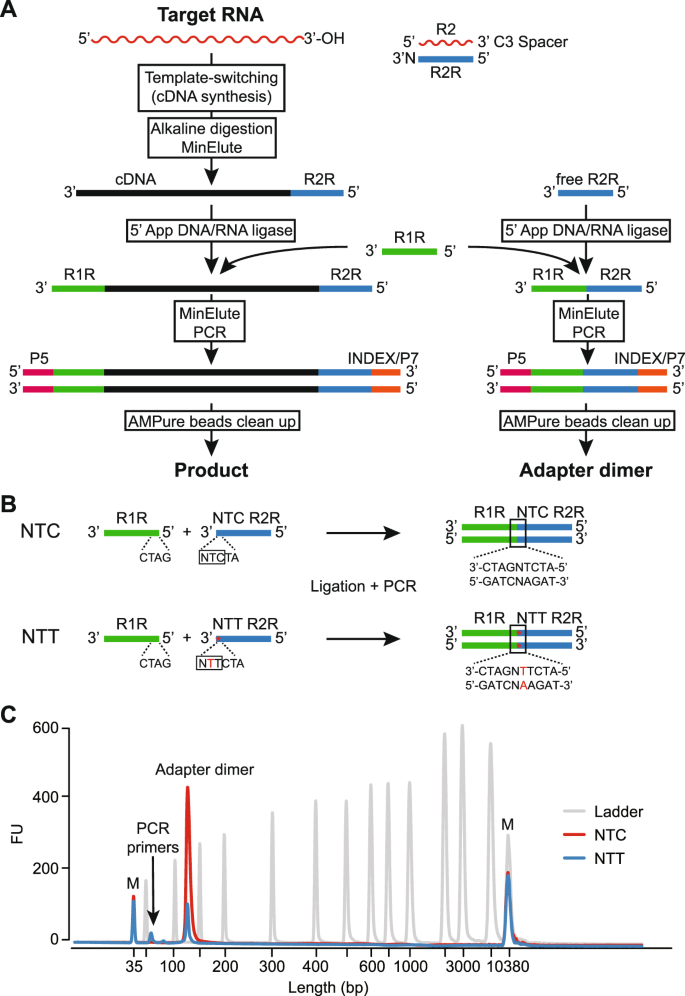
Improved TGIRT-seq methods for comprehensive transcriptome profiling with decreased adapter dimer formation and bias correction | Scientific Reports

High-throughput, Efficient, and Unbiased Capture of Small RNAs from Low-input Samples for Sequencing | Scientific Reports

Figure 1 from Quantitative Bias in Illumina TruSeq and a Novel Post Amplification Barcoding Strategy for Multiplexed DNA and Small RNA Deep Sequencing | Semantic Scholar
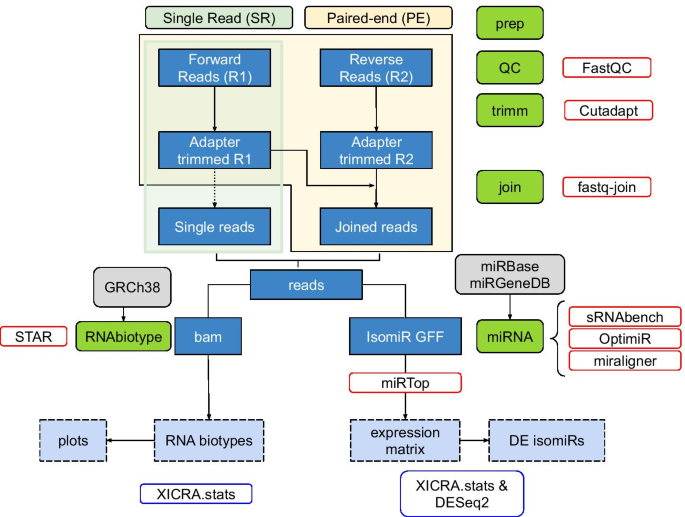
Paired-end small RNA sequencing reveals a possible overestimation in the isomiR sequence repertoire previously reported from conventional single read data analysis | BMC Bioinformatics | Full Text