IJMS | Free Full-Text | T-Cell Receptor Repertoire Sequencing and Its Applications: Focus on Infectious Diseases and Cancer
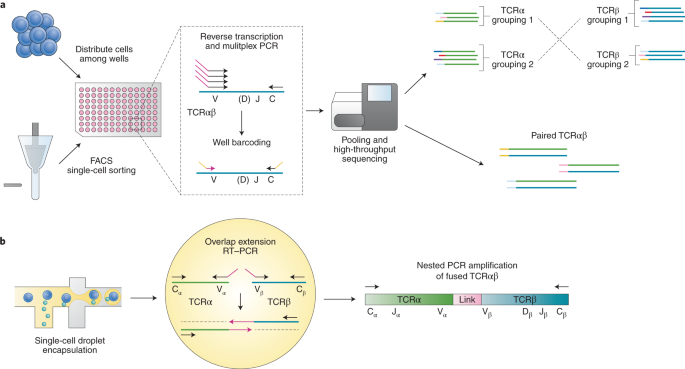
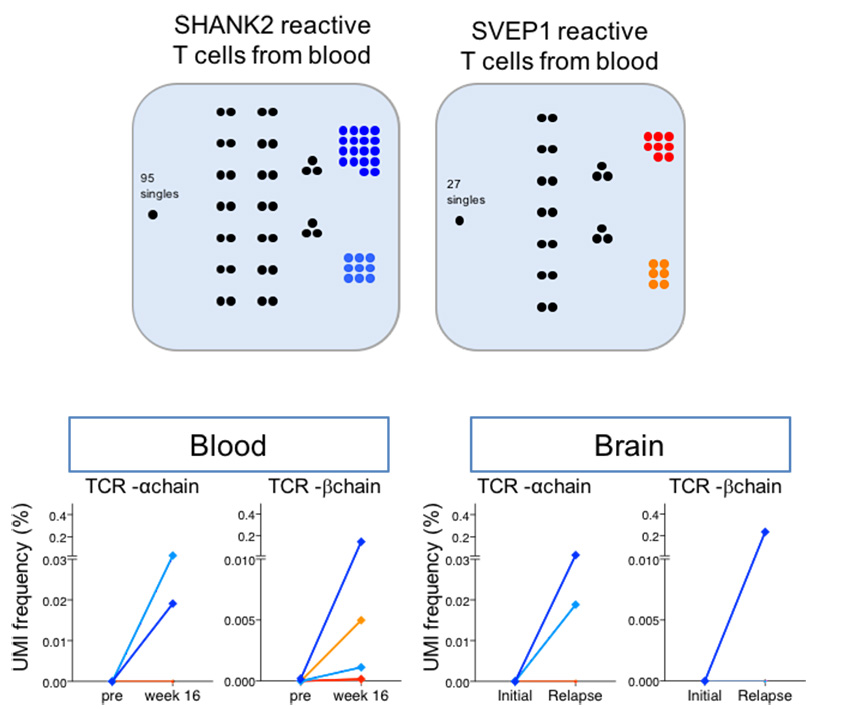
Single-cell RNA sequencing coupled to TCR profiling of large granular lymphocyte leukemia T cells | Nature Communications

Frontiers | Quantitative Characterization of the T Cell Receptor Repertoire of Naïve and Memory Subsets Using an Integrated Experimental and Computational Pipeline Which Is Robust, Economical, and Versatile
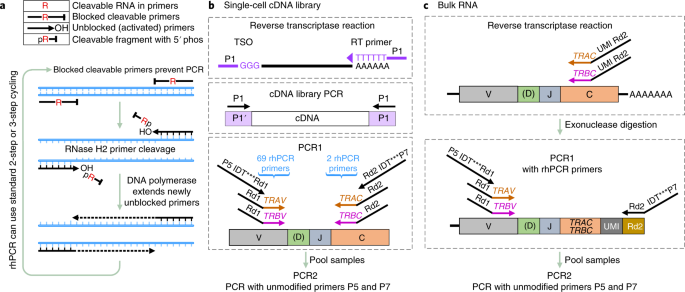
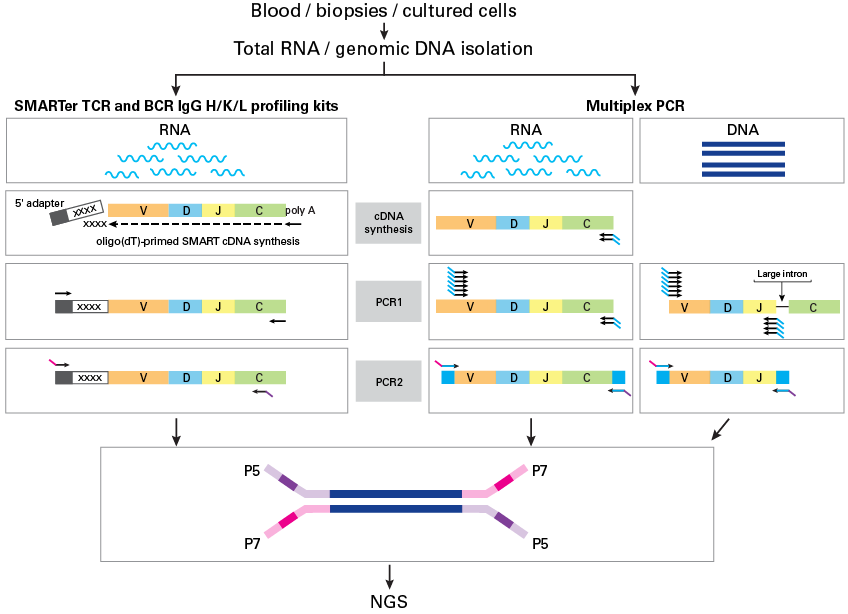
RNase H–dependent PCR-enabled T-cell receptor sequencing for highly specific and efficient targeted sequencing of T-cell receptor mRNA for single-cell and repertoire analysis | Nature Protocols

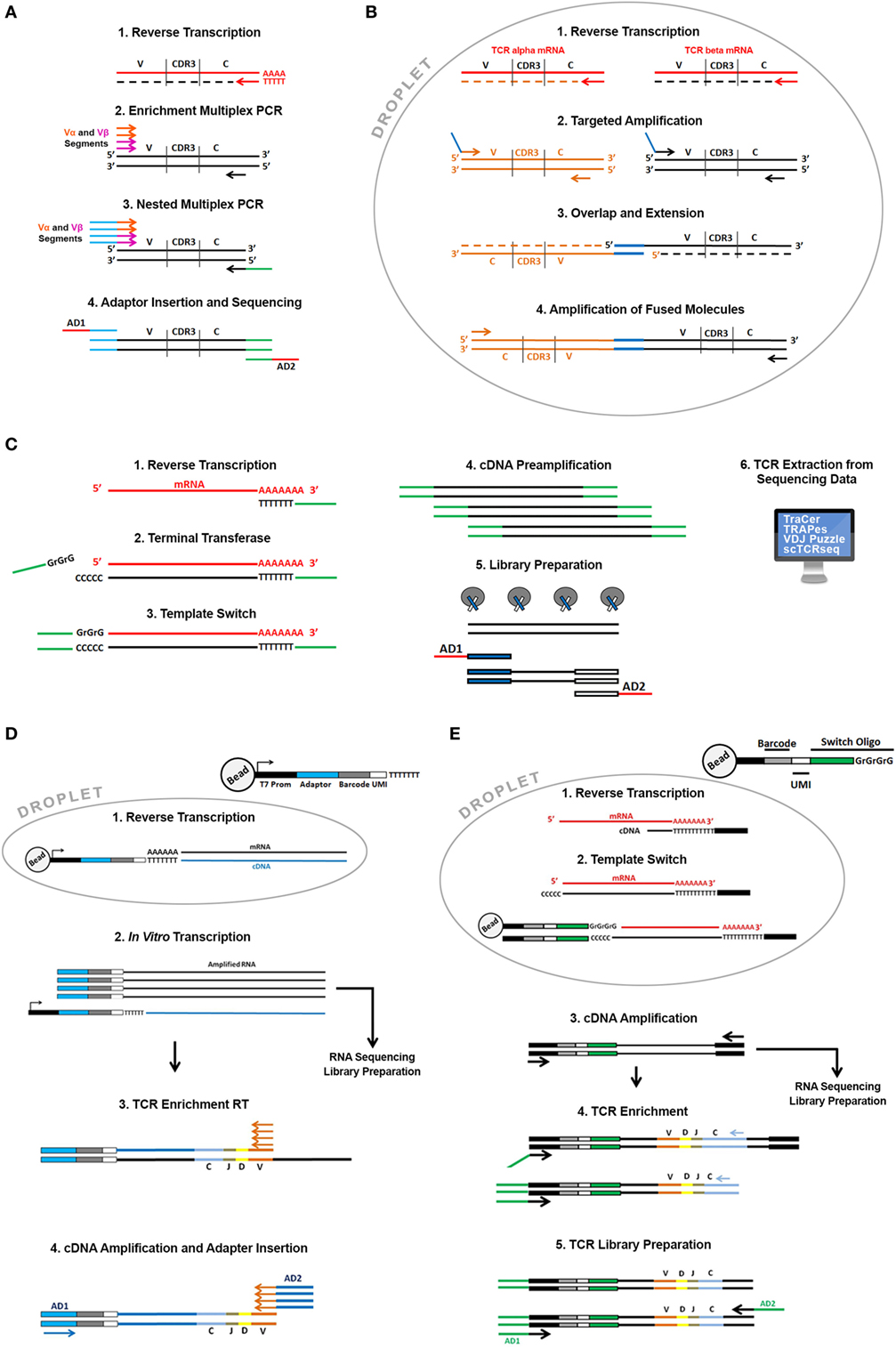
Schematic of the target TCR molecule in Protocols A and B showing the... | Download Scientific Diagram

Ultra-efficient sequencing of T Cell receptor repertoires reveals shared responses in muscle from patients with Myositis - eBioMedicine
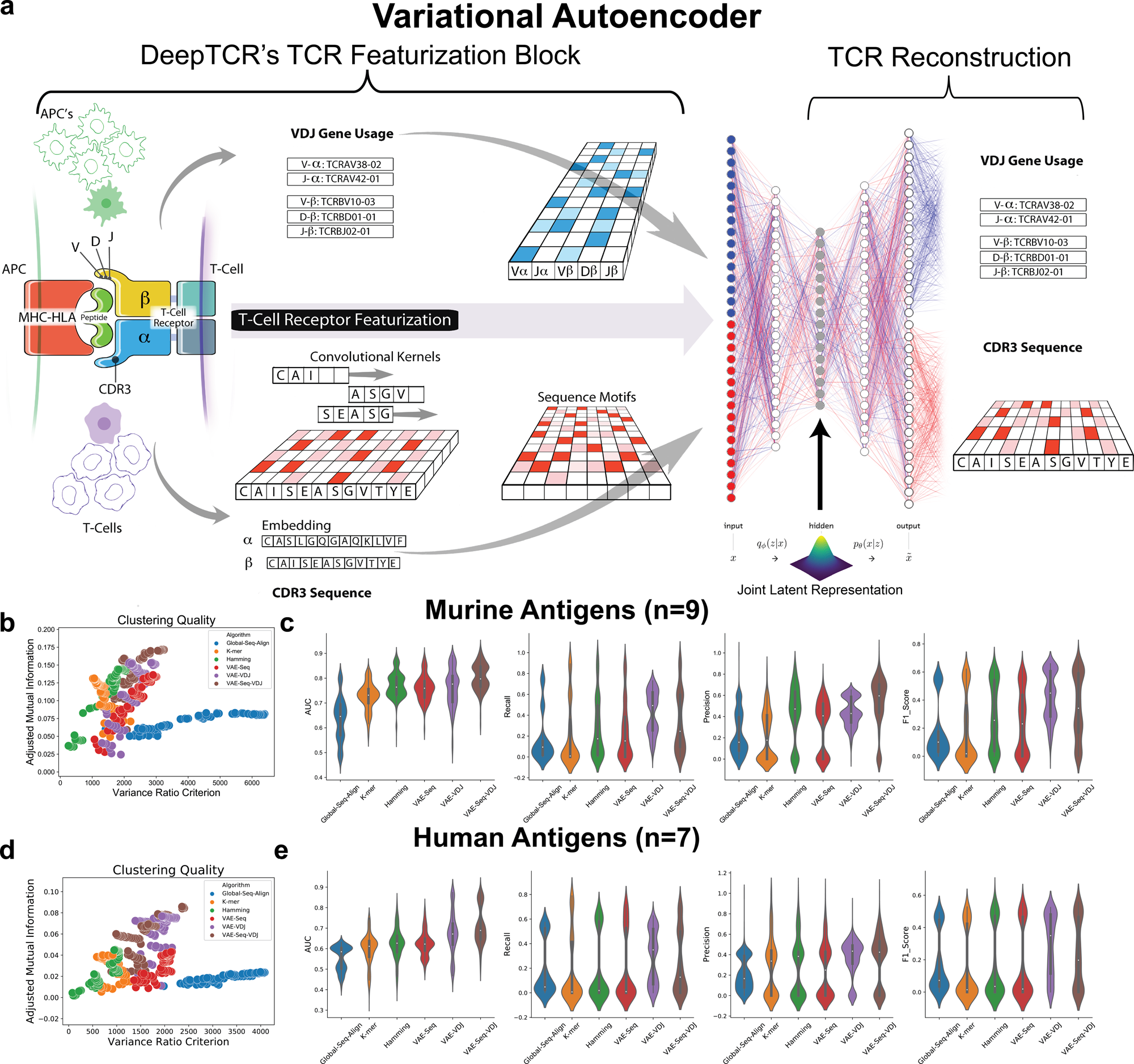
DeepTCR is a deep learning framework for revealing sequence concepts within T-cell repertoires | Nature Communications
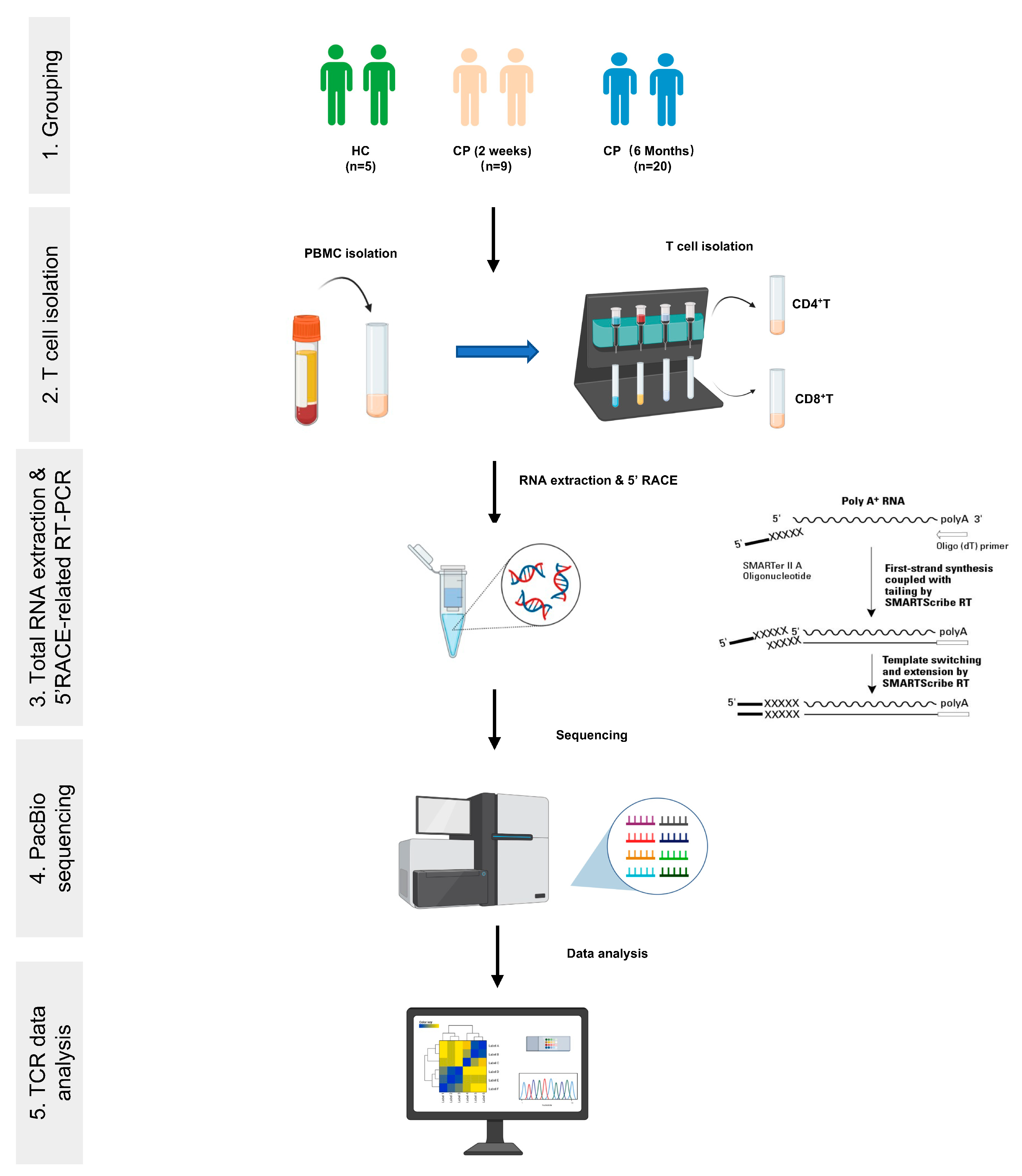
Cells | Free Full-Text | Analysis of TCR Repertoire by High-Throughput Sequencing Indicates the Feature of T Cell Immune Response after SARS-CoV-2 Infection

Rapid cloning, expression, and functional characterization of paired αβ and γδ T-cell receptor chains from single-cell analysis: Molecular Therapy - Methods & Clinical Development
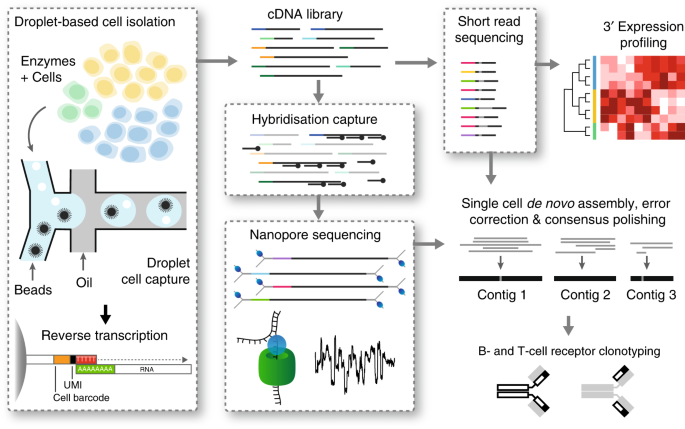
High-throughput targeted long-read single cell sequencing reveals the clonal and transcriptional landscape of lymphocytes | Nature Communications