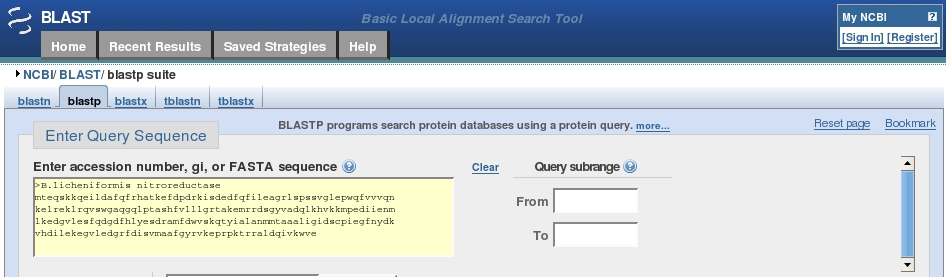
Steps to create non-redundant patent sequence databases at two levels.... | Download Scientific Diagram
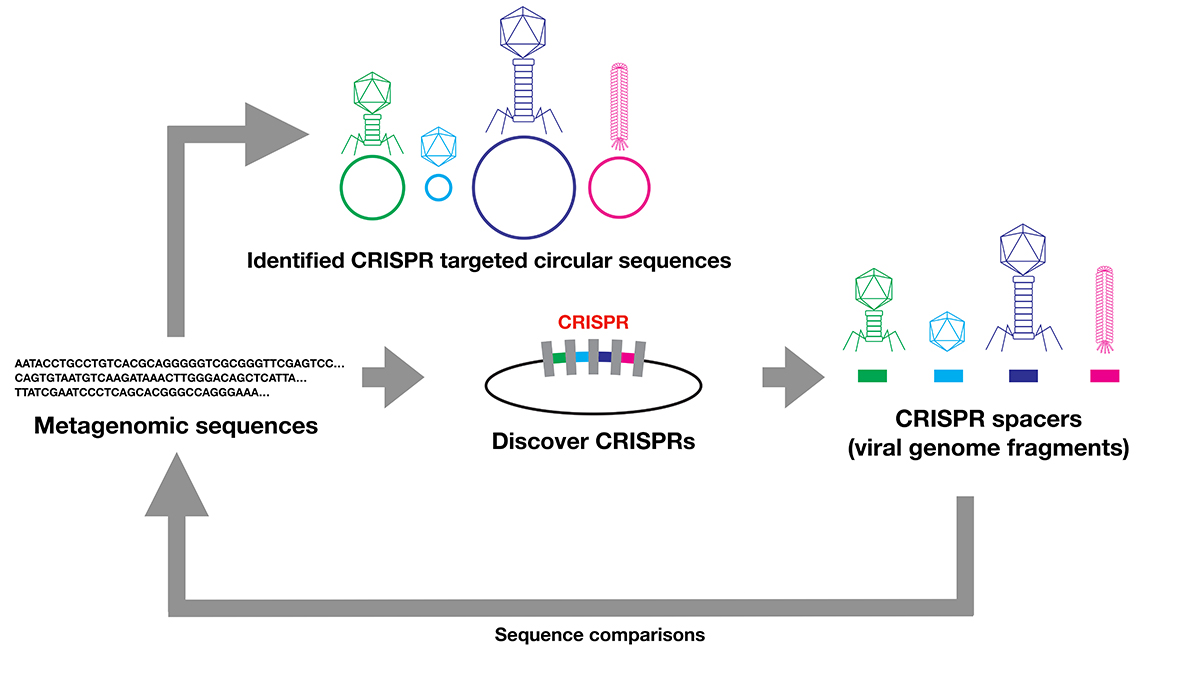
Comprehensive discovery of CRISPR-targeted terminally redundant sequences in the human gut metagenome: viruses, plasmids, and more::National Institute of Genetics

PROTEIN DATABASES. The ideal sequence database for computational analyses and data-mining: I t must be complete with minimal redundancy It must contain. - ppt download

Drop on flow should remove redundant incoming and outgoing sequence flows. · Issue #774 · bpmn-io/bpmn-js · GitHub
Comprehensive discovery of CRISPR-targeted terminally redundant sequences in the human gut metagenome: Viruses, plasmids, and more | PLOS Computational Biology
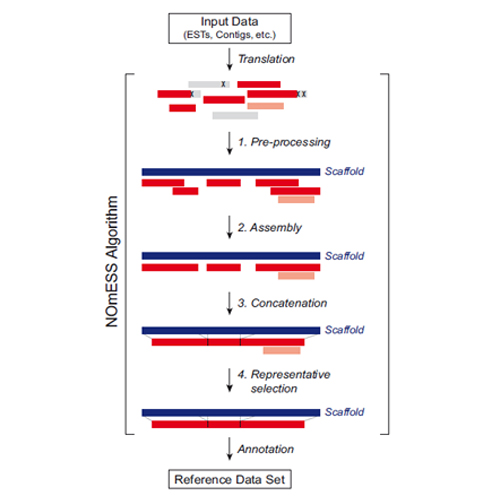
CIPSM - Homology-driven assembly of NOn-redundant protEin sequence sets (NOmESS) for mass spectrometry
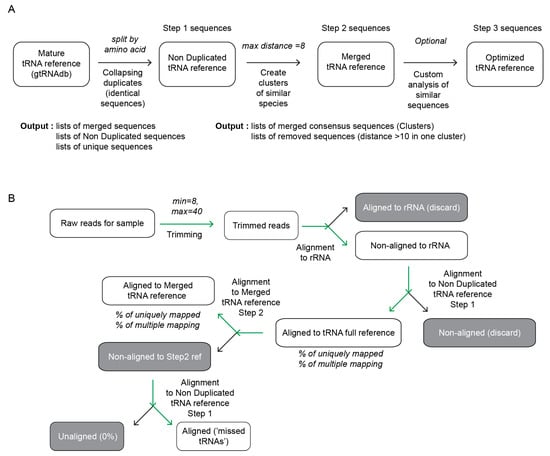
Genes | Free Full-Text | Non-Redundant tRNA Reference Sequences for Deep Sequencing Analysis of tRNA Abundance and Epitranscriptomic RNA Modifications

An open-sourced bioinformatic pipeline for the processing of Next-Generation Sequencing derived nucleotide reads: Identification and authentication of ancient metagenomic DNA | bioRxiv

Building the non-redundant sequence set. The schema depicts an example... | Download Scientific Diagram

MGnify on Twitter: "A new version of the proteinDB has been released on @MGnifyDB. More than double in size, it now has over 2.4 billion non-redundant sequences comprising 623 million clusters. Internal



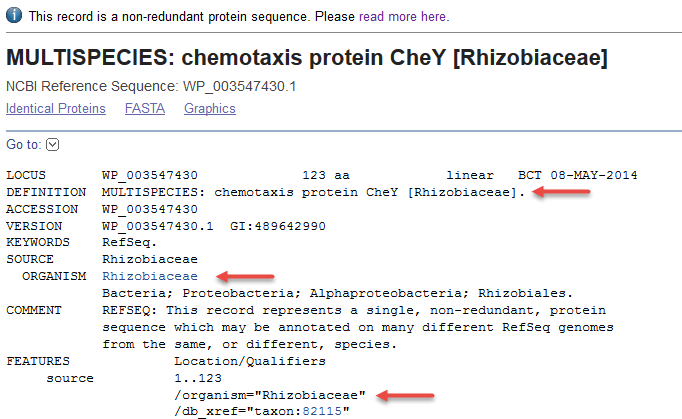

![PDF] Comprehensive Analysis of Non Redundant Protein Database | Semantic Scholar PDF] Comprehensive Analysis of Non Redundant Protein Database | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/e08e0cec115b9c9d85ff0f0ab0bd0f1a4e7e7f26/5-Figure1-1.png)