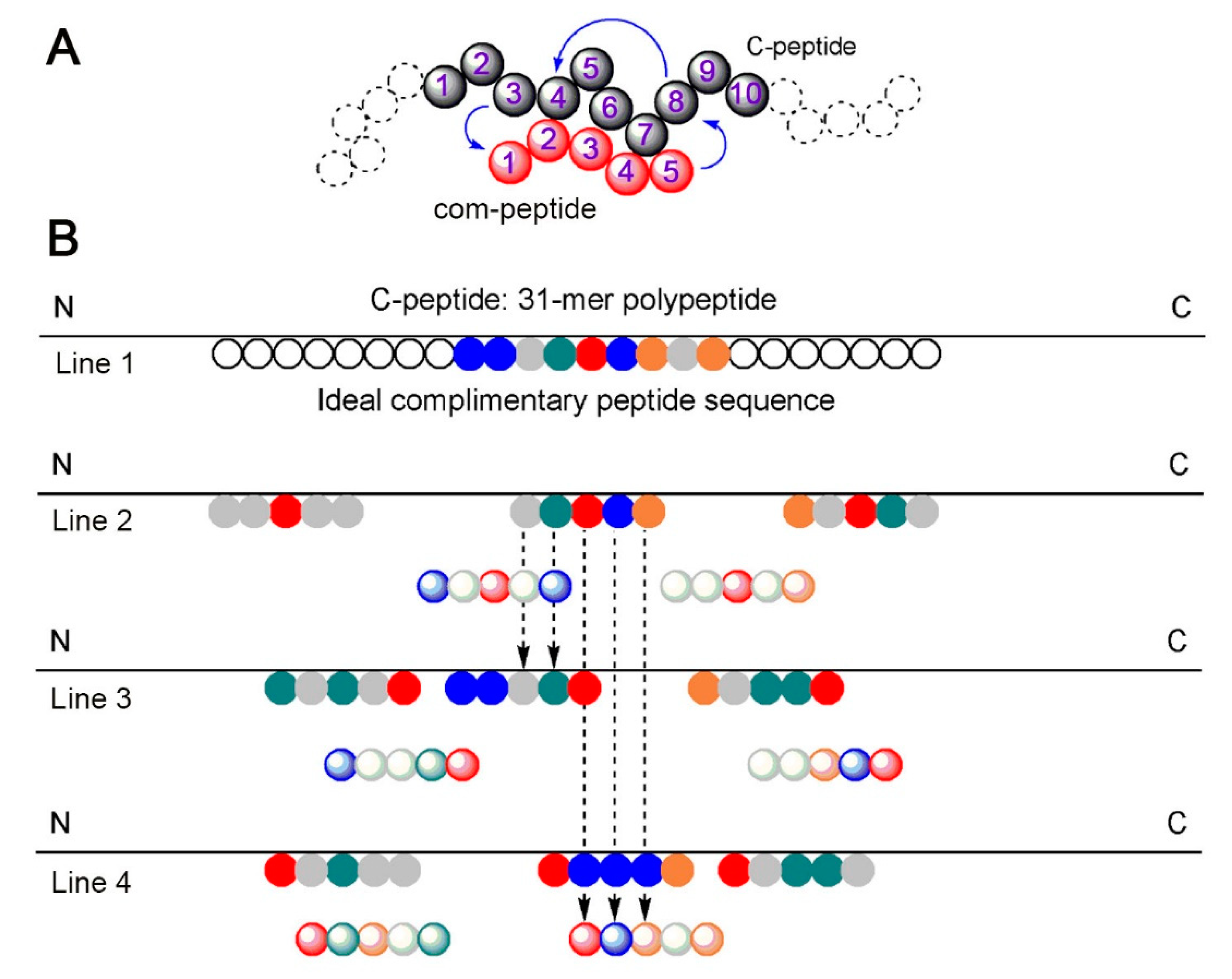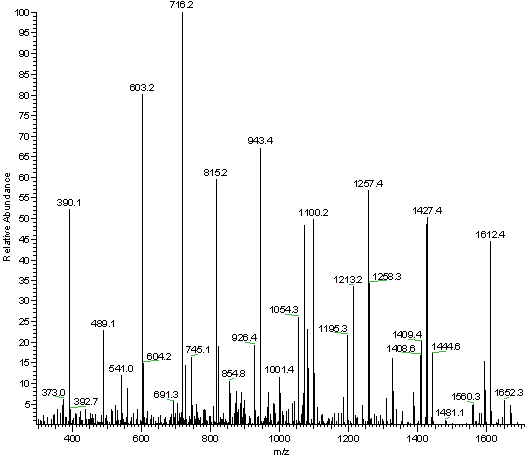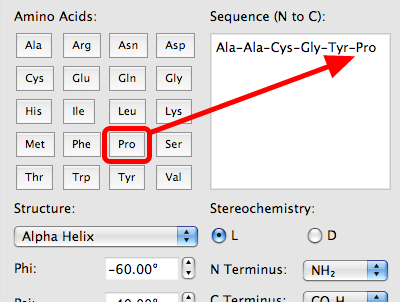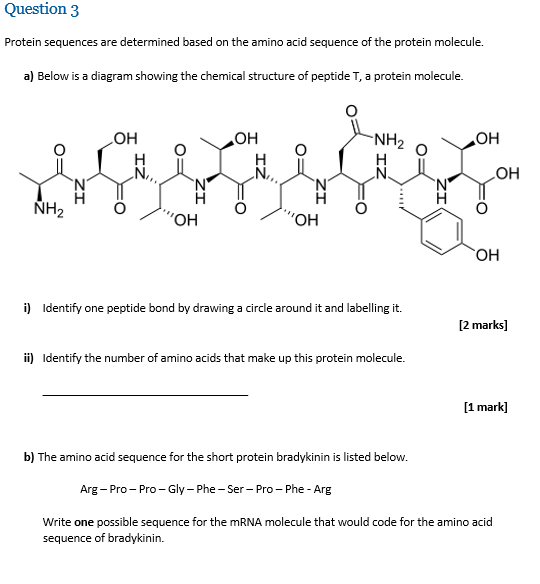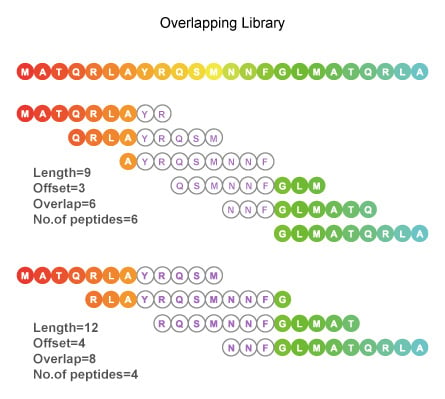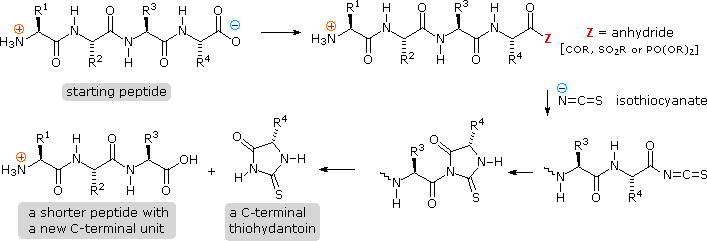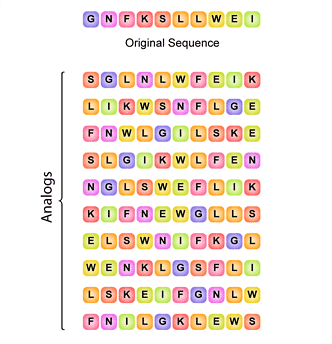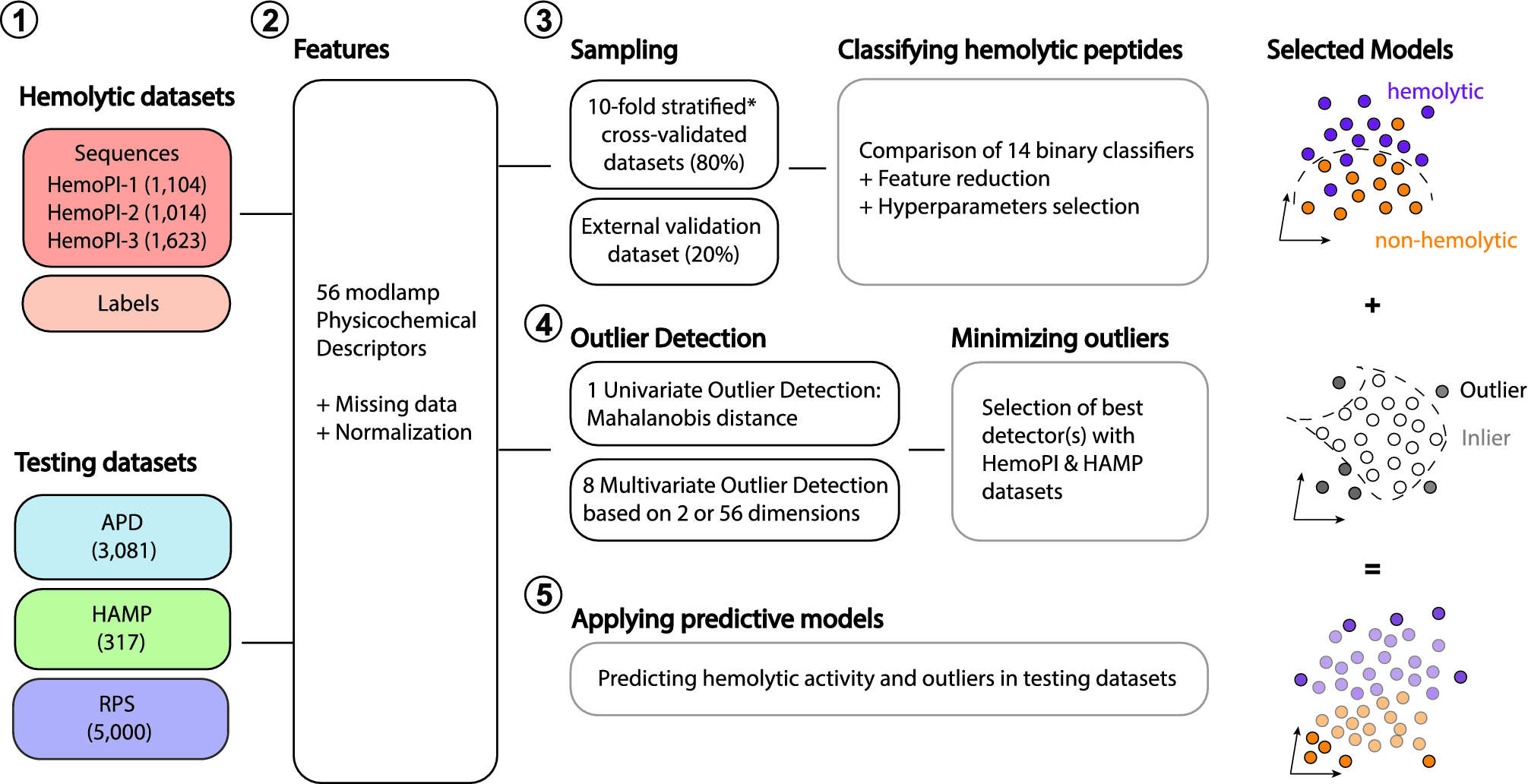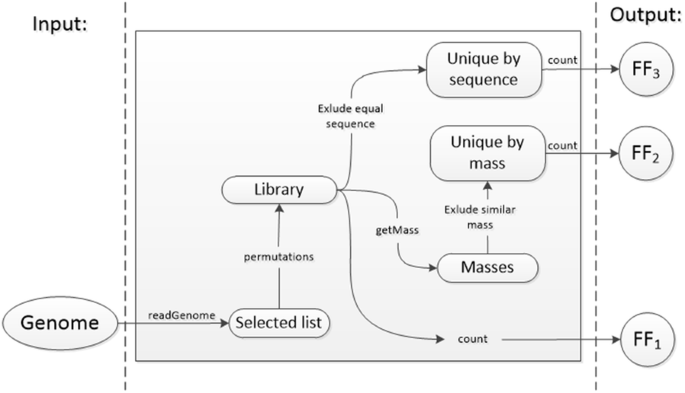
Algorithm-supported, mass and sequence diversity-oriented random peptide library design | Journal of Cheminformatics | Full Text

PepSeA: Peptide Sequence Alignment and Visualization Tools to Enable Lead Optimization | Journal of Chemical Information and Modeling
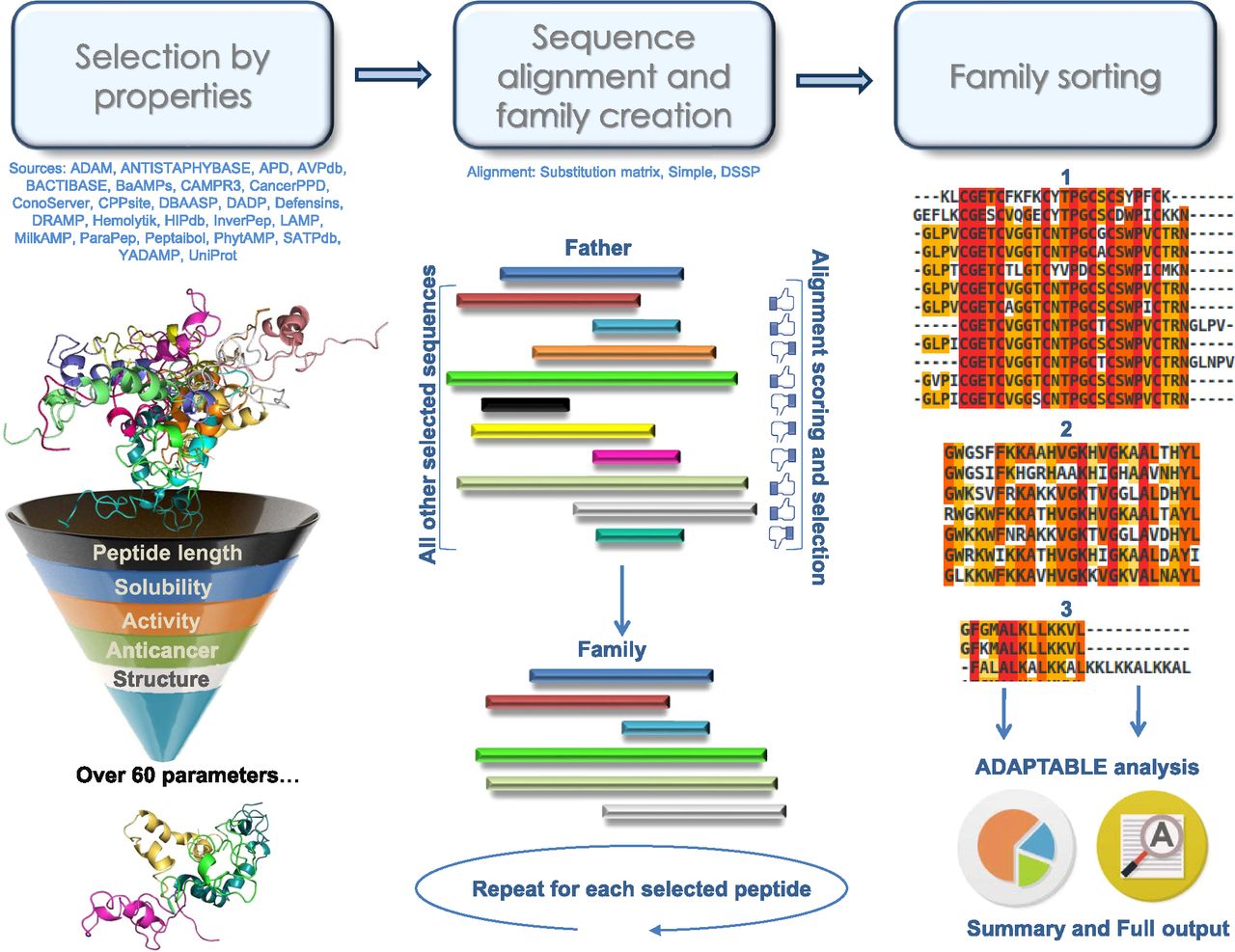
ADAPTABLE: a comprehensive web platform of antimicrobial peptides tailored to the user's research | Life Science Alliance

ADAPTABLE: a comprehensive web platform of antimicrobial peptides tailored to the user's research | Life Science Alliance
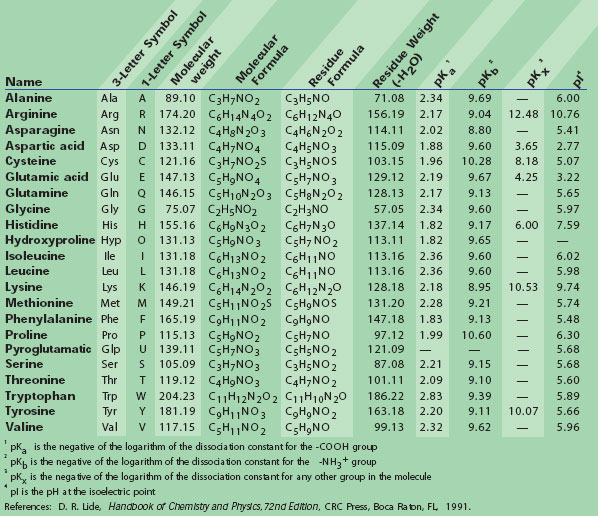
![PDF] PseKRAAC: a flexible web server for generating pseudo K-tuple reduced amino acids composition | Semantic Scholar PDF] PseKRAAC: a flexible web server for generating pseudo K-tuple reduced amino acids composition | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/bde9663b6cac60d4380e3827c0257604b29a8527/2-Table1-1.png)
PDF] PseKRAAC: a flexible web server for generating pseudo K-tuple reduced amino acids composition | Semantic Scholar

PepSeA: Peptide Sequence Alignment and Visualization Tools to Enable Lead Optimization | Journal of Chemical Information and Modeling
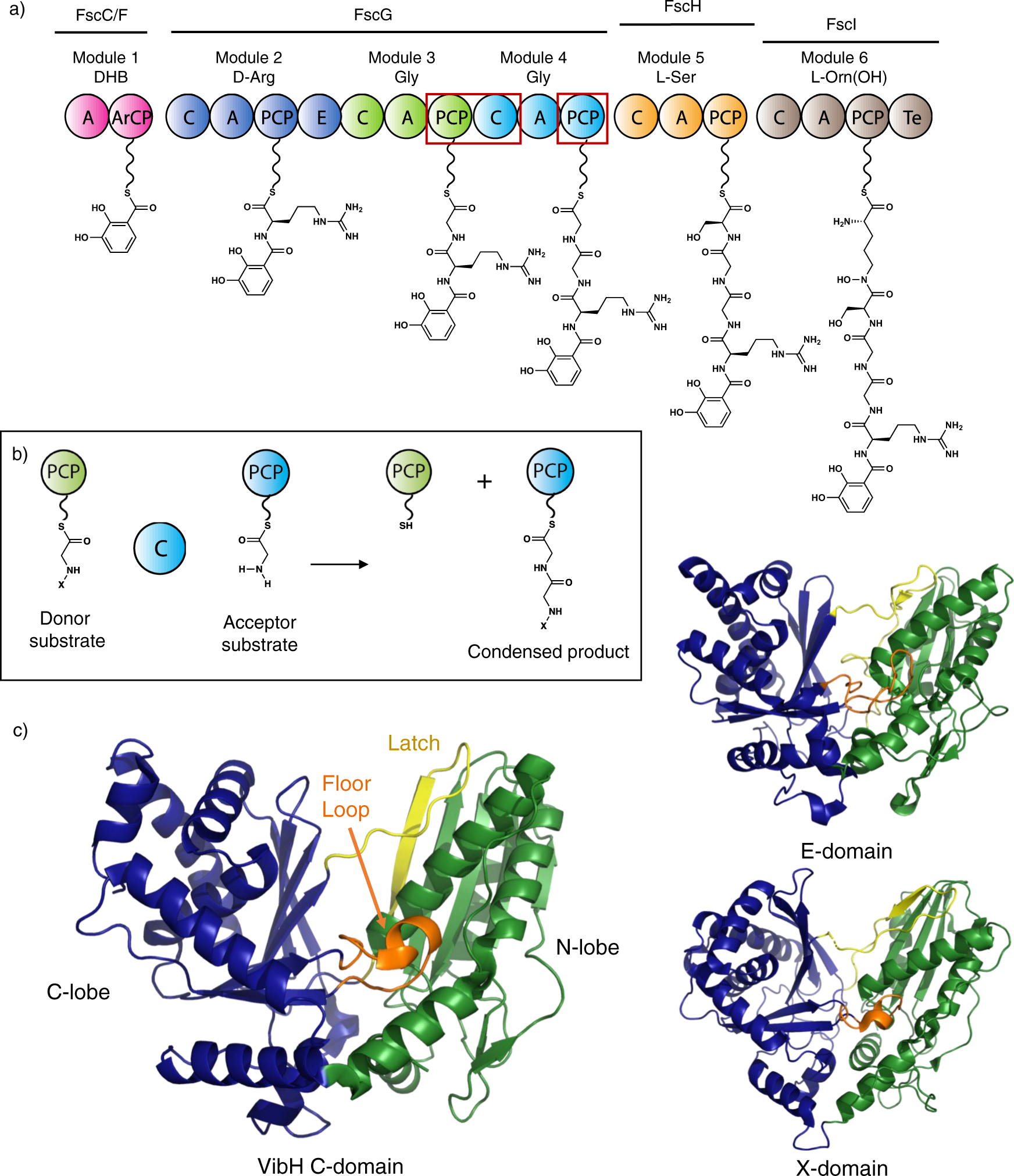
Structures of a non-ribosomal peptide synthetase condensation domain suggest the basis of substrate selectivity | Nature Communications

Uncovering Thousands of New Peptides with Sequence-Mask-Search Hybrid De Novo Peptide Sequencing Framework - ScienceDirect
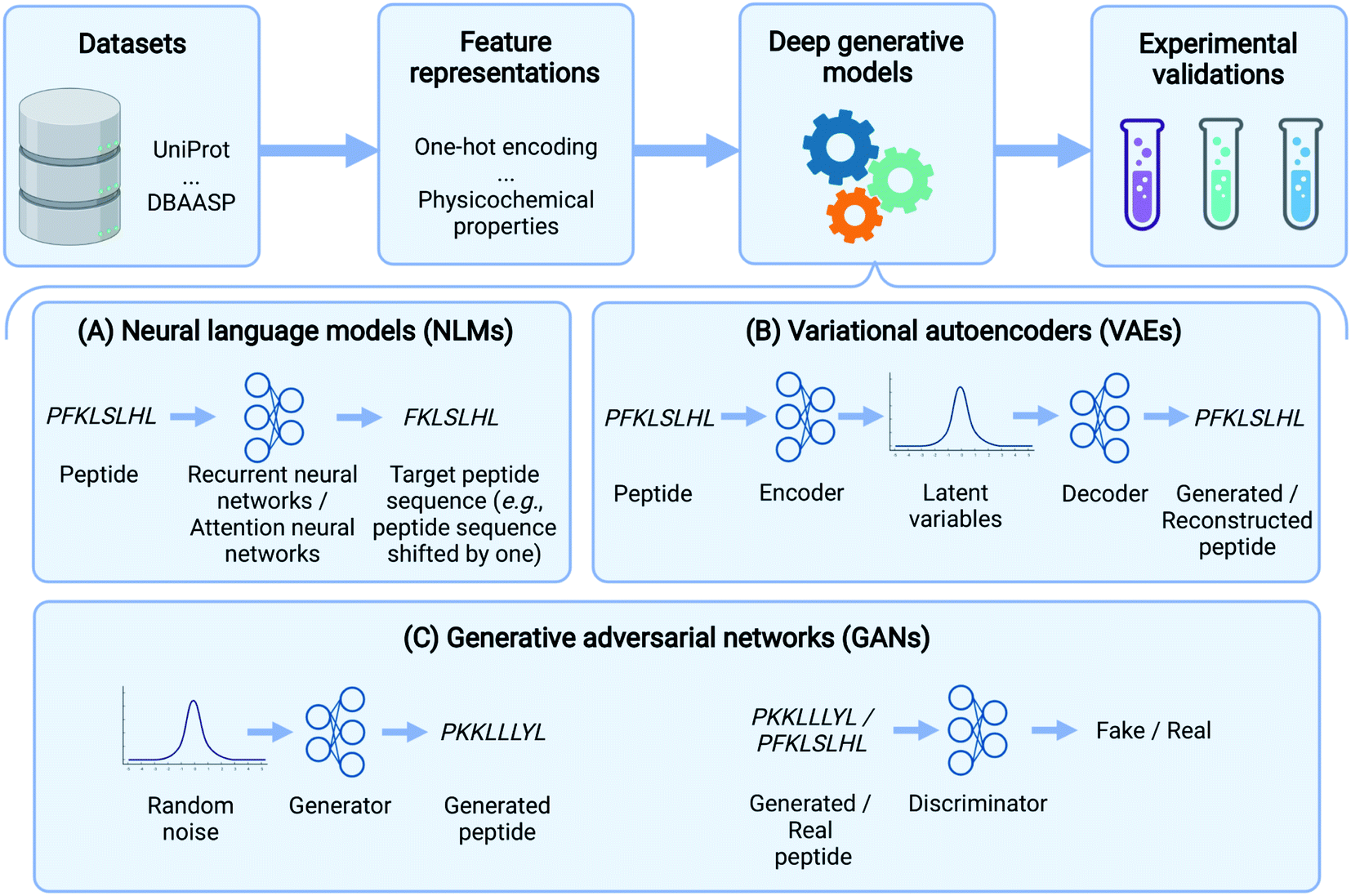
Deep generative models for peptide design - Digital Discovery (RSC Publishing) DOI:10.1039/D1DD00024A