Identification of functional PAM sequences for CRISPR-interference.... | Download Scientific Diagram
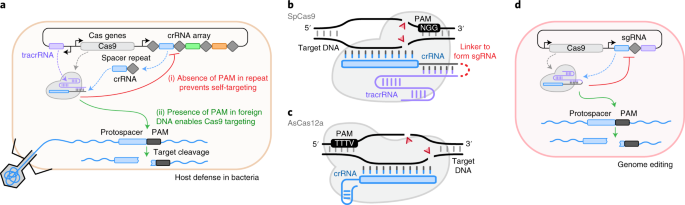
Scalable characterization of the PAM requirements of CRISPR–Cas enzymes using HT-PAMDA | Nature Protocols
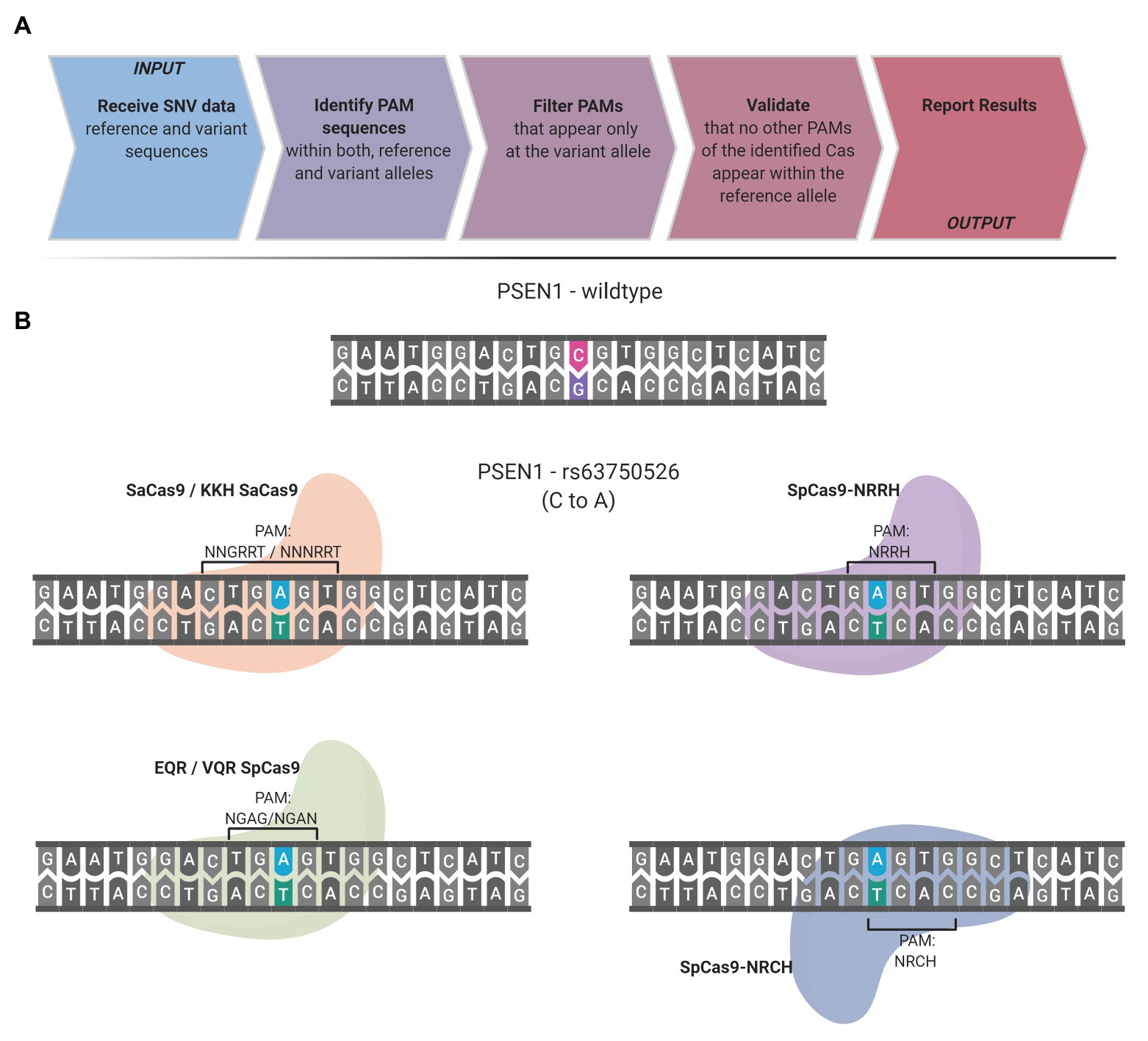
Frontiers | CrisPam: SNP-Derived PAM Analysis Tool for Allele-Specific Targeting of Genetic Variants Using CRISPR-Cas Systems