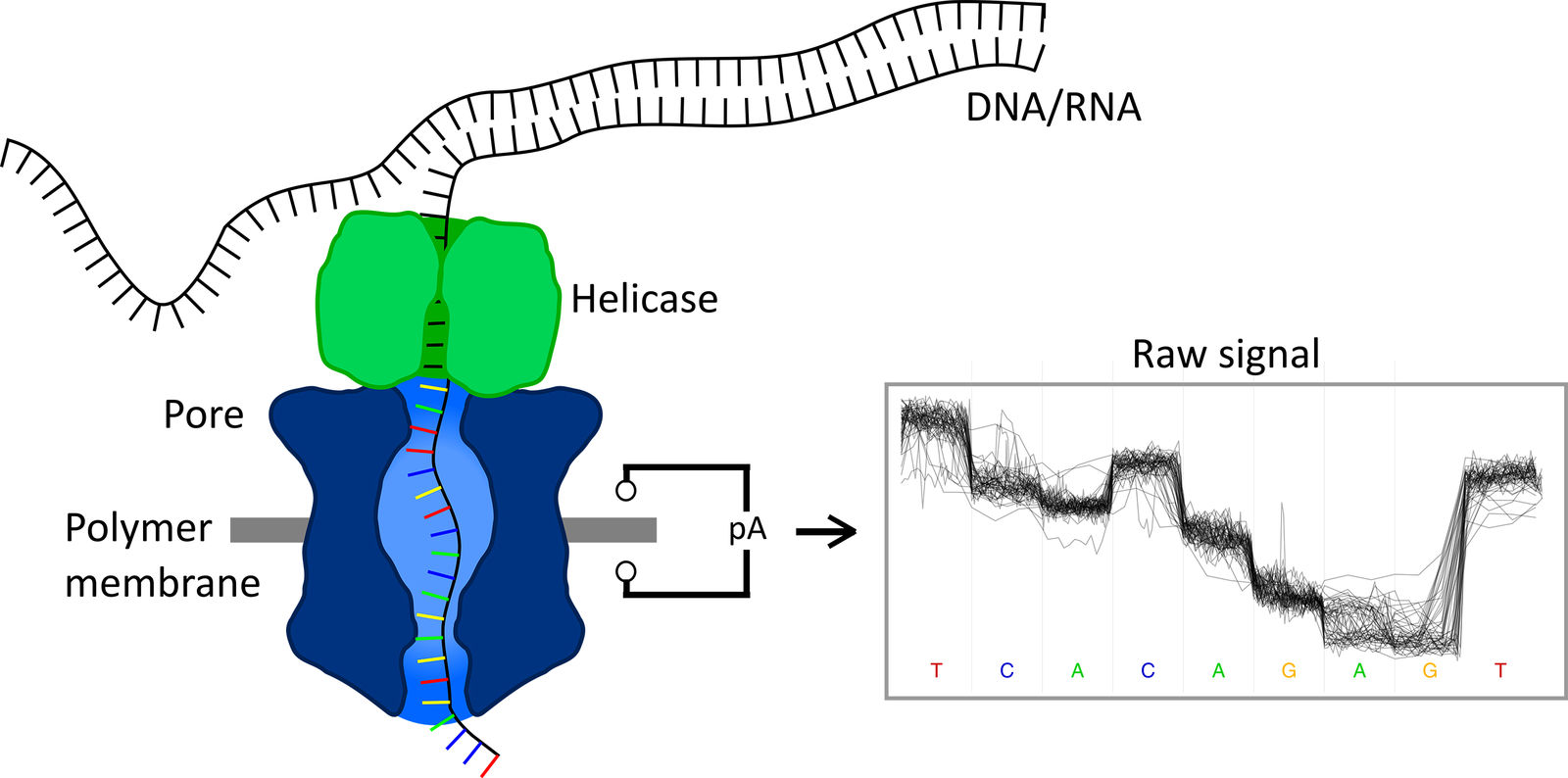
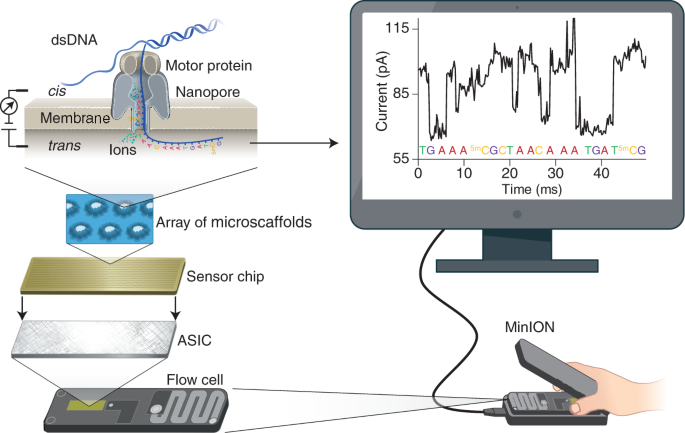
Applications and potentials of nanopore sequencing in the (epi)genome and (epi)transcriptome era - ScienceDirect

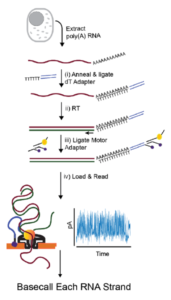
Nanopore-based native RNA sequencing provides insights into prokaryotic transcription, operon structures, rRNA maturation and modifications | bioRxiv
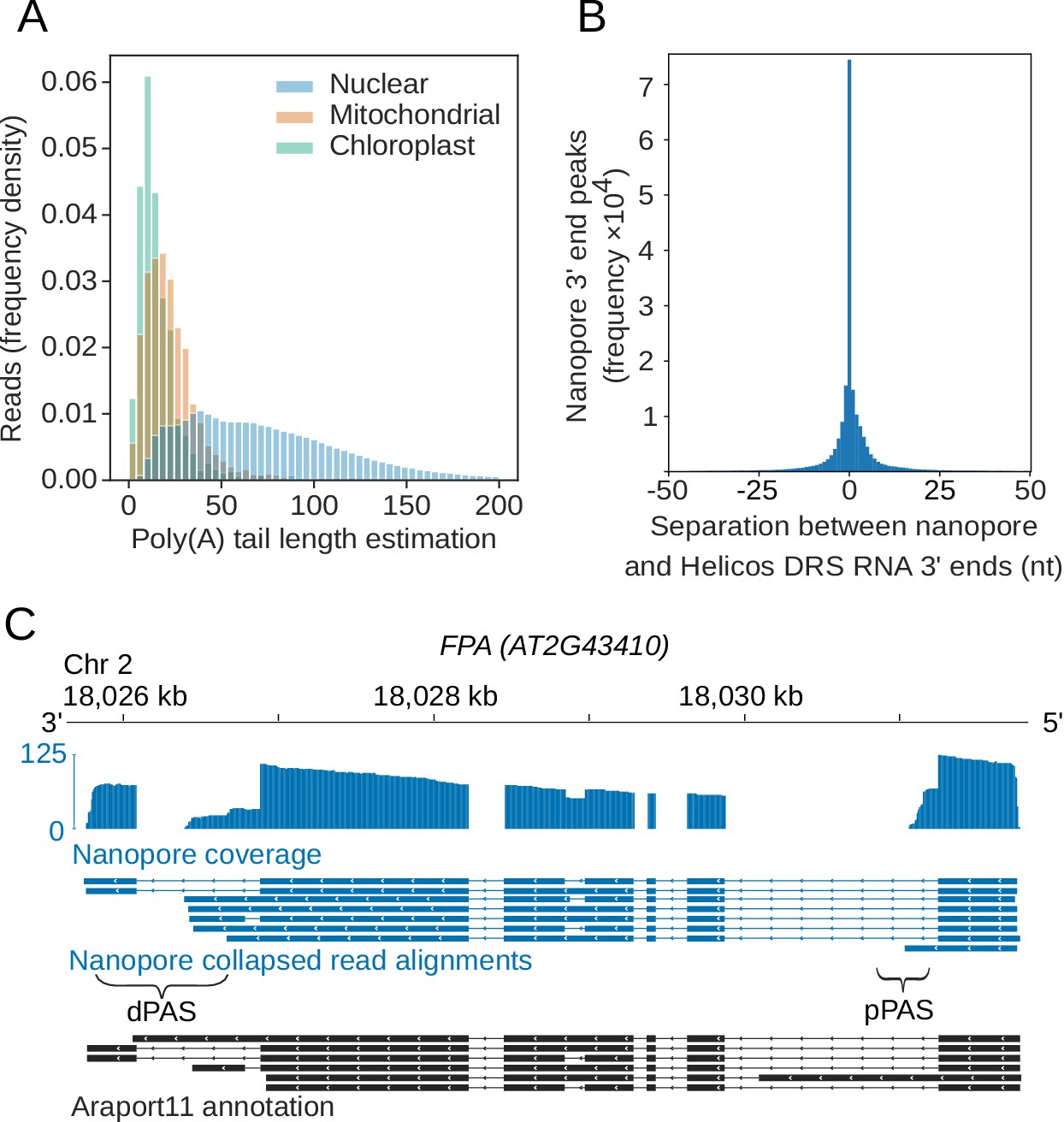
Nanopore direct RNA sequencing maps the complexity of Arabidopsis mRNA processing and m6A modification | eLife

Direct detection of RNA modifications and structure using single molecule nanopore sequencing | bioRxiv
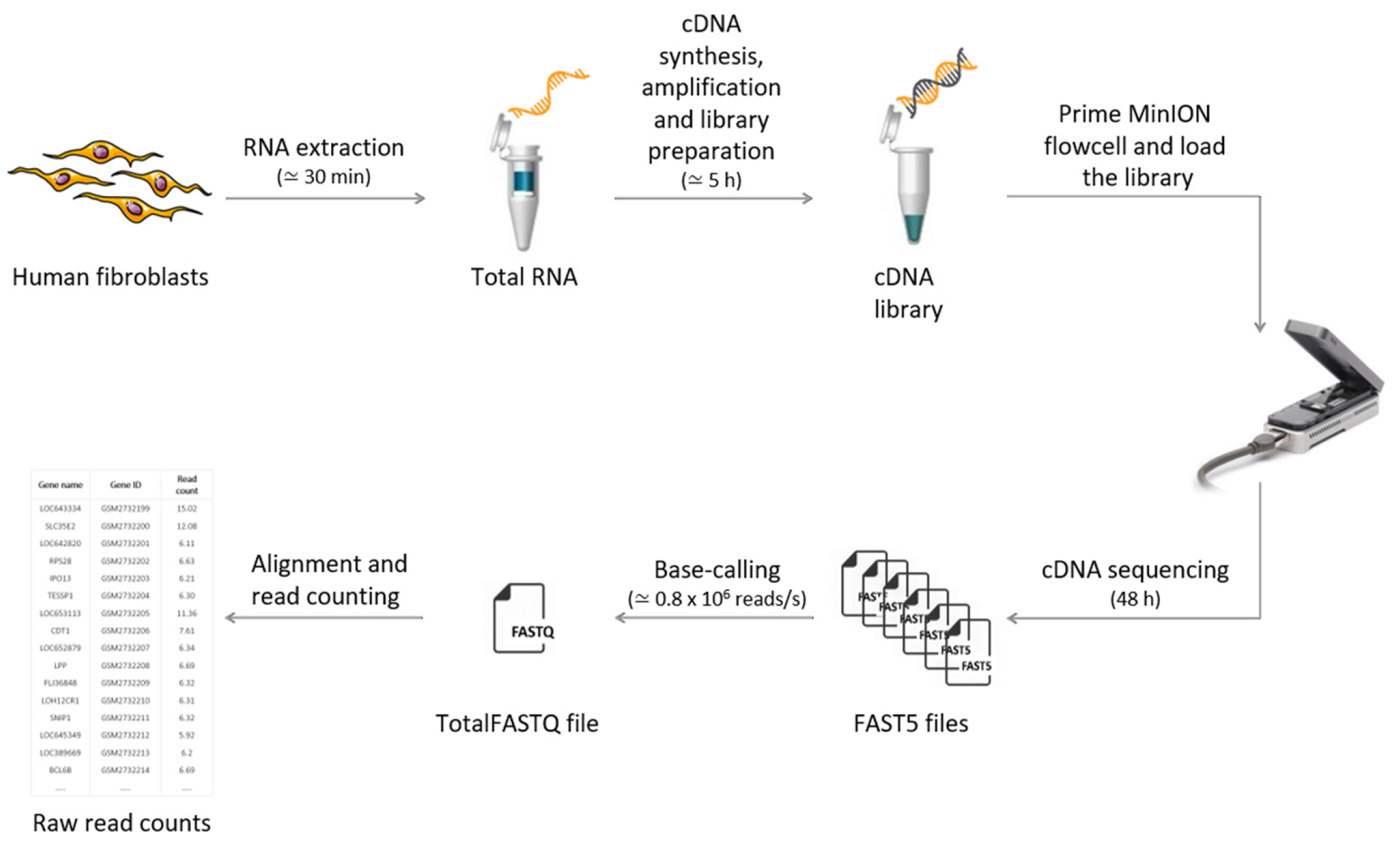
IJMS | Free Full-Text | Evaluation of Oxford Nanopore MinION RNA-Seq Performance for Human Primary Cells

RNA on Twitter: "This review provides an overview on direct RNA nanopore sequencing technologies, and how it can be used to detect chemical modifications in native RNAs https://t.co/B5VFj0Nzwt https://t.co/sCb4SxuVdi" / Twitter
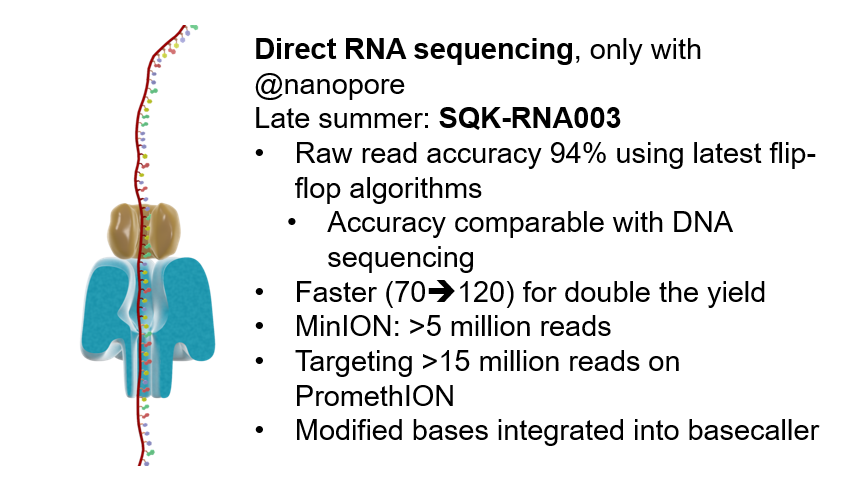

Oxford Nanopore on Twitter: "AH: Direct RNA with @nanopore will get closer to mainstream method this year #nanoporeconf https://t.co/3LrUy8fGog" / Twitter

PORE-cupine – determination of isoform-specific RNA structure with nanopore long reads | RNA-Seq Blog