
Tina Han on Twitter: "RT @razoralign: Enabling high-accuracy long-read amplicon sequences using unique molecular identifiers with Nanopore or PacBio sequencing h…" / Twitter
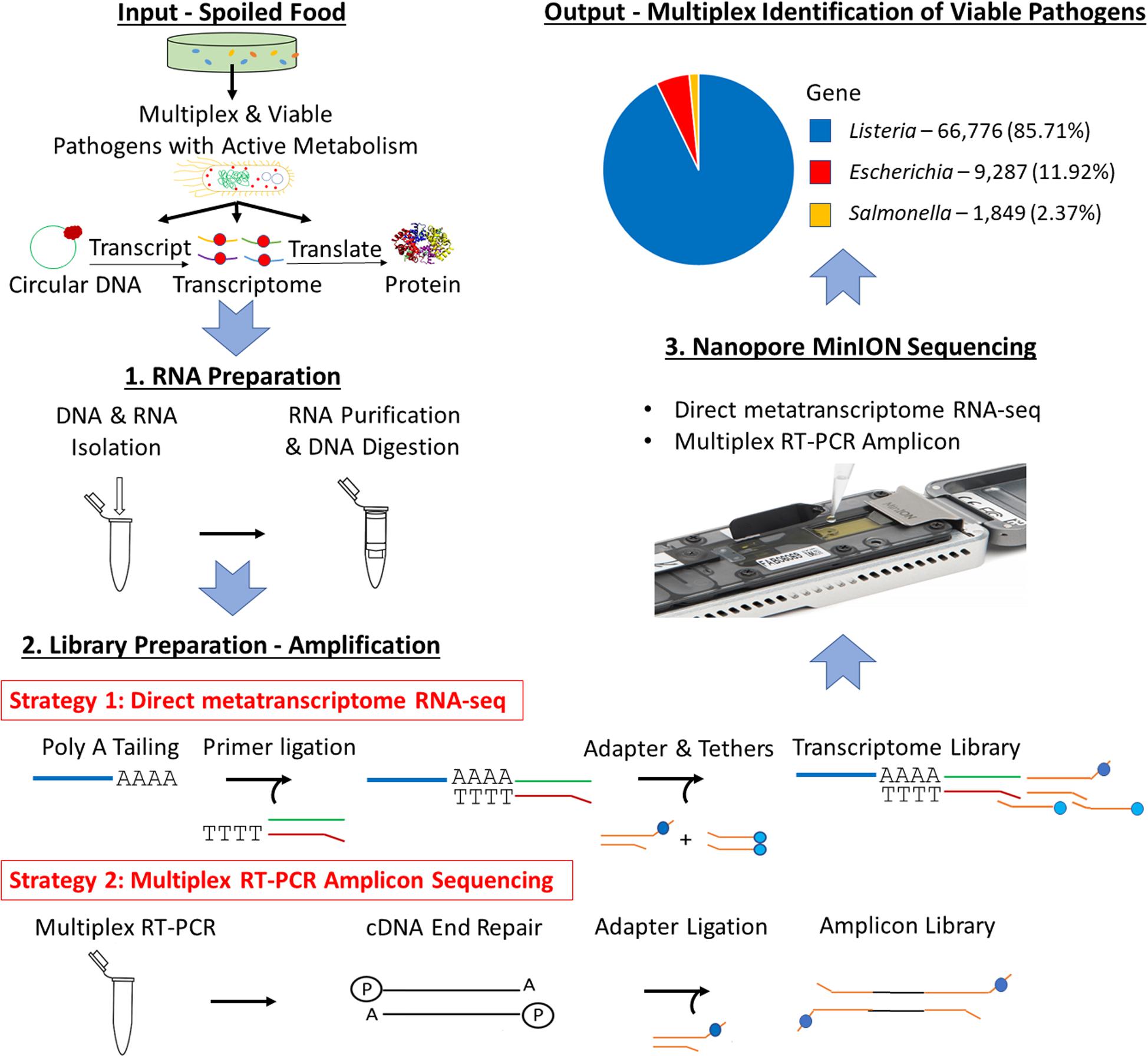
Frontiers | Direct Metatranscriptome RNA-seq and Multiplex RT-PCR Amplicon Sequencing on Nanopore MinION – Promising Strategies for Multiplex Identification of Viable Pathogens in Food
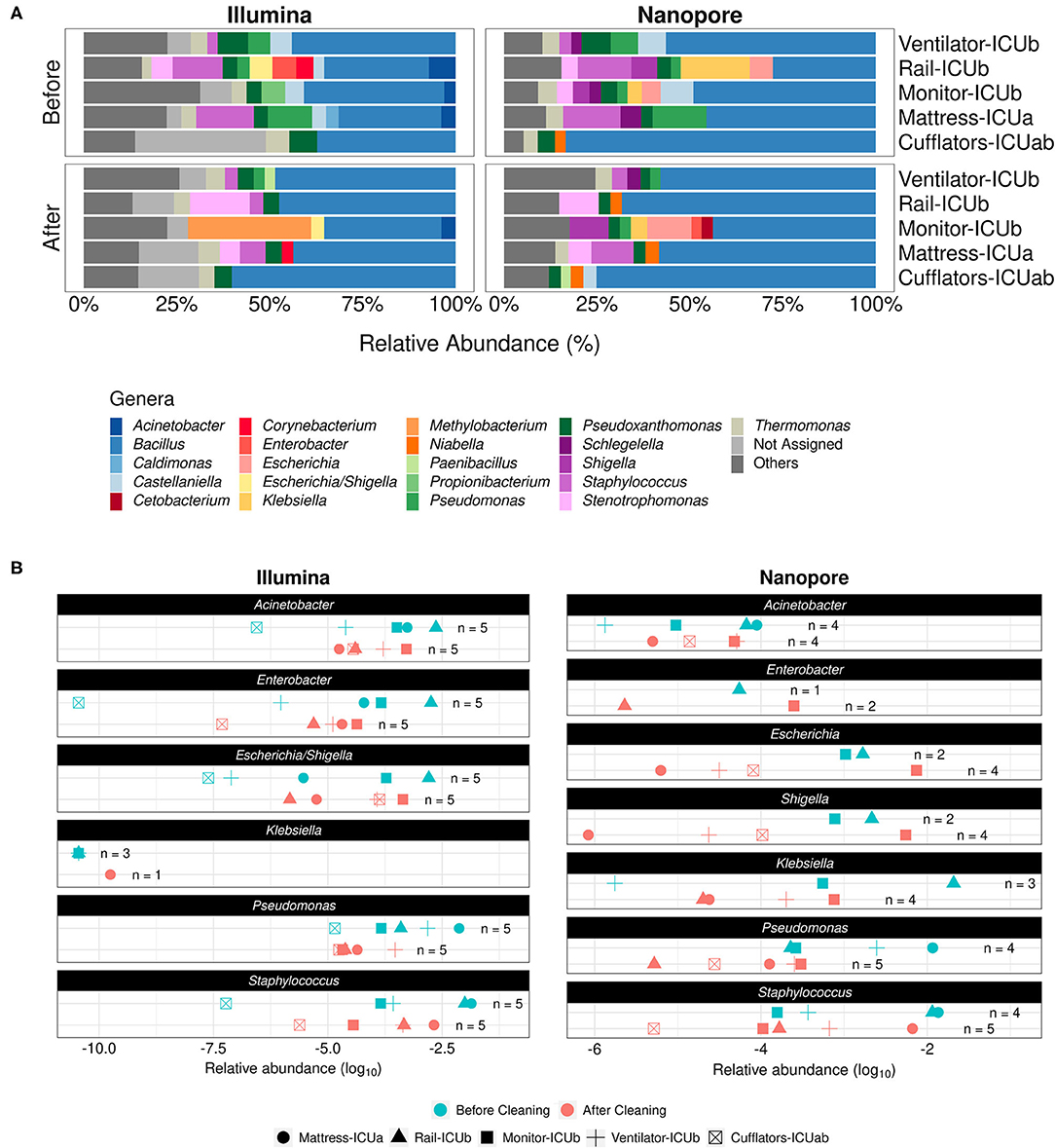
Frontiers | Nanopore Sequencing Provides Rapid and Reliable Insight Into Microbial Profiles of Intensive Care Units

NanoRTax, a real-time pipeline for taxonomic and diversity analysis of nanopore 16S rRNA amplicon sequencing data - ScienceDirect
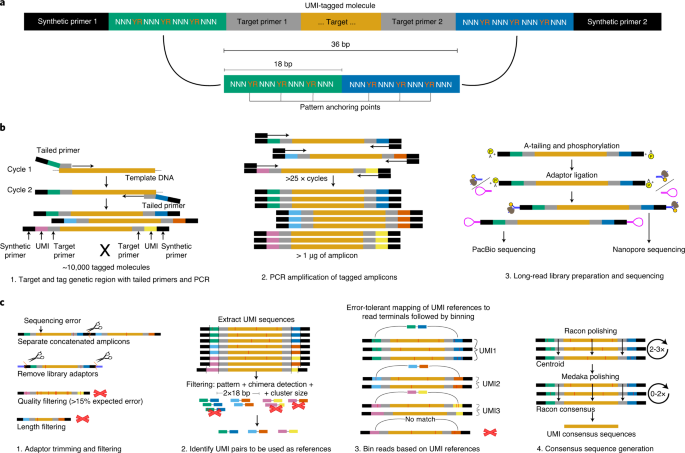
High-accuracy long-read amplicon sequences using unique molecular identifiers with Nanopore or PacBio sequencing | Nature Methods

Oncogene Concatenated Enriched Amplicon Nanopore Sequencing for rapid, accurate, and affordable somatic mutation detection | Genome Biology | Full Text
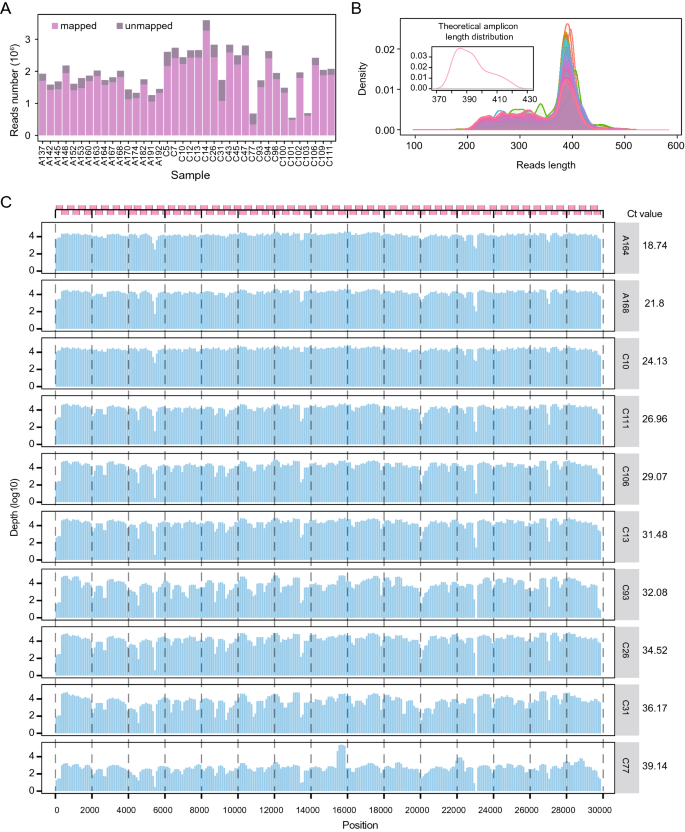
Rapid Acquisition of High-Quality SARS-CoV-2 Genome via Amplicon-Oxford Nanopore Sequencing | SpringerLink

High-accuracy long-read amplicon sequences using unique molecular identifiers with Nanopore or PacBio sequencing | Nature Methods
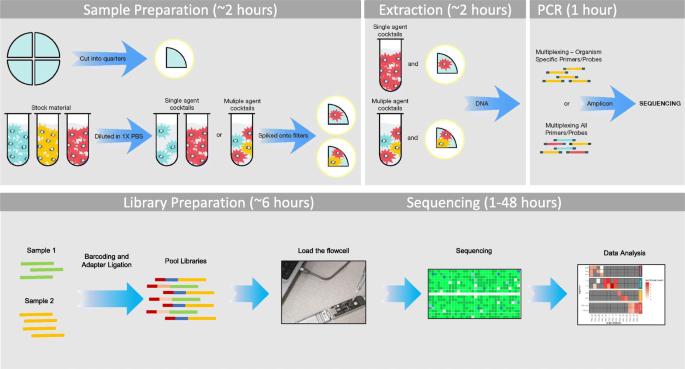
Comparison of the performance of an amplicon sequencing assay based on Oxford Nanopore technology to real-time PCR assays for detecting bacterial biodefense pathogens | BMC Genomics | Full Text
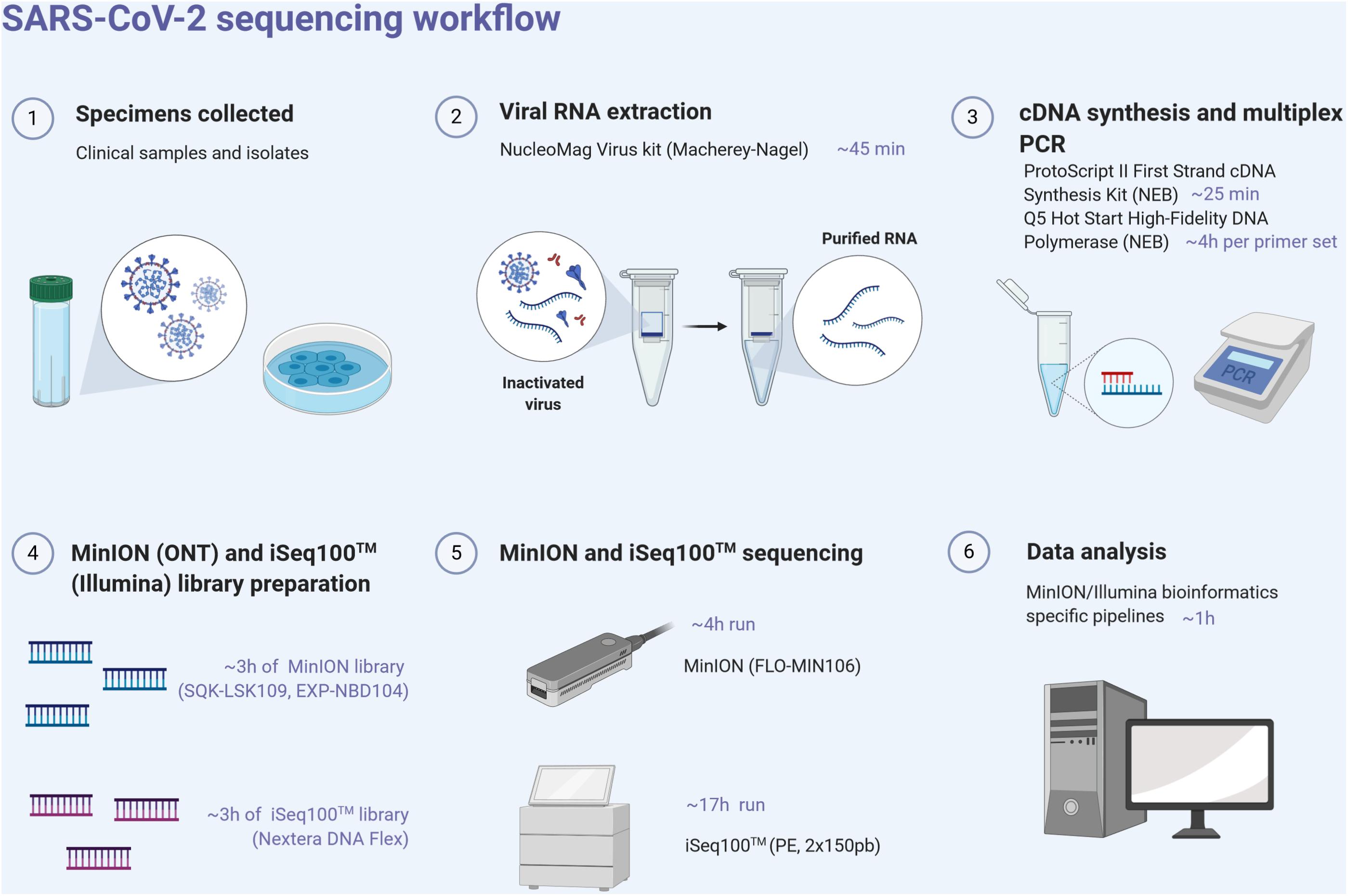
Frontiers | Rapid Genomic Characterization of SARS-CoV-2 by Direct Amplicon-Based Sequencing Through Comparison of MinION and Illumina iSeq100TM System
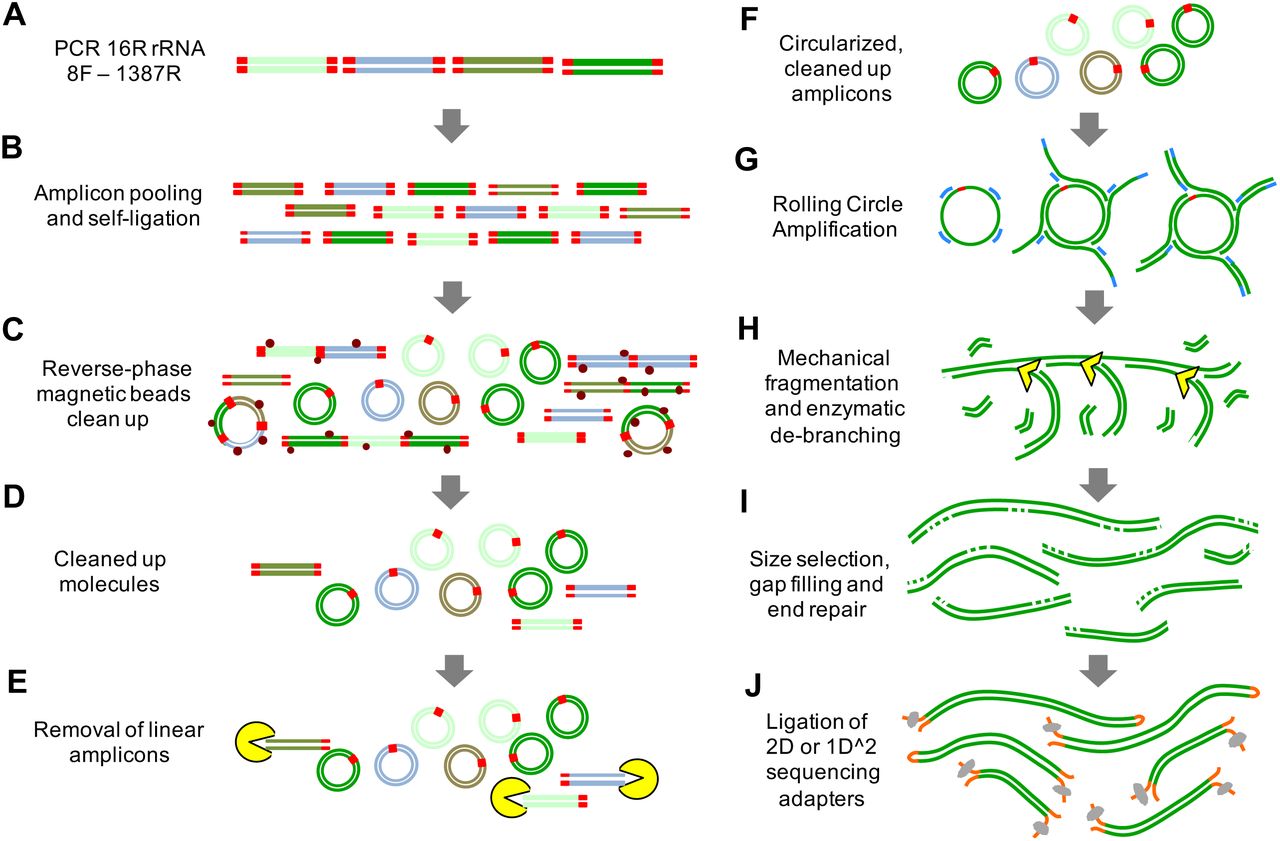
NanoAmpli-Seq: A workflow for amplicon sequencing from mixed microbial communities on the nanopore sequencing platform | bioRxiv
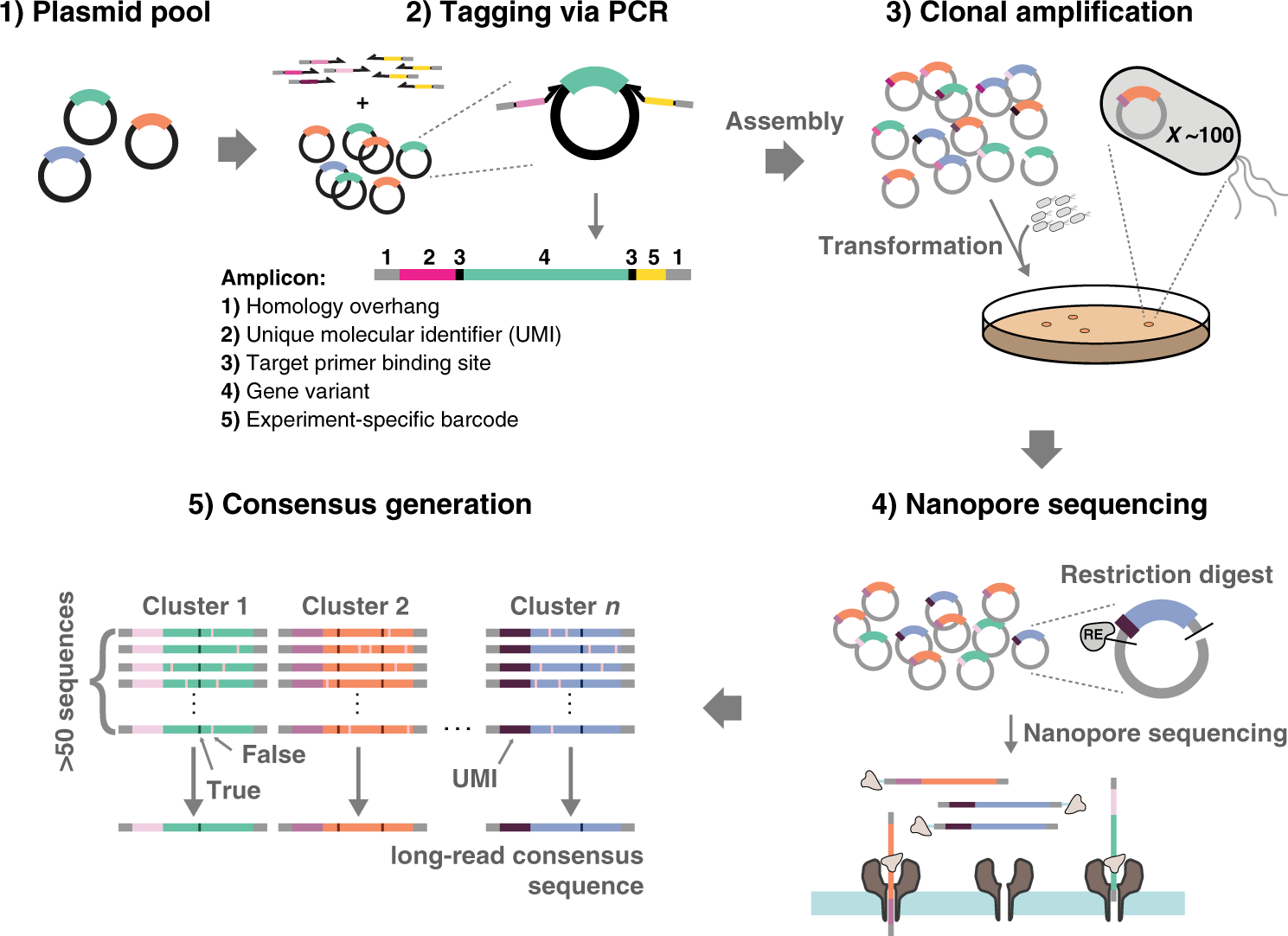
UMI-linked consensus sequencing enables phylogenetic analysis of directed evolution | Nature Communications
![Evaluation of full-length nanopore 16S sequencing for detection of pathogens in microbial keratitis [PeerJ] Evaluation of full-length nanopore 16S sequencing for detection of pathogens in microbial keratitis [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2021/10778/1/fig-1-full.png)
Evaluation of full-length nanopore 16S sequencing for detection of pathogens in microbial keratitis [PeerJ]

NanoAmpli-Seq: A workflow for amplicon sequencing from mixed microbial communities on the nanopore sequencing platform
Concatenation of short amplicons by Stochastic Amplicon Ligation (SAL).... | Download Scientific Diagram

Proof of concept for multiplex amplicon sequencing for mutation identification using the MinION nanopore sequencer | Scientific Reports

Microorganisms | Free Full-Text | Simplified Point-of-Care Full SARS-CoV-2 Genome Sequencing Using Nanopore Technology
Boxplot of sequencing depth data and amplicons size (bp). The range of... | Download Scientific Diagram
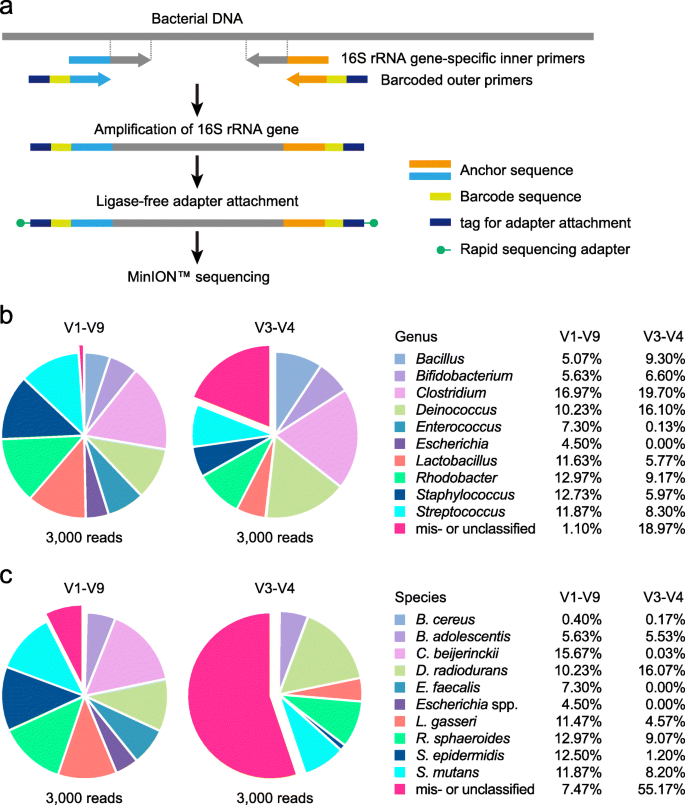





![De novo species identification using 16S rRNA gene nanopore sequencing [PeerJ] De novo species identification using 16S rRNA gene nanopore sequencing [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/10029/1/fig-1-full.png)

