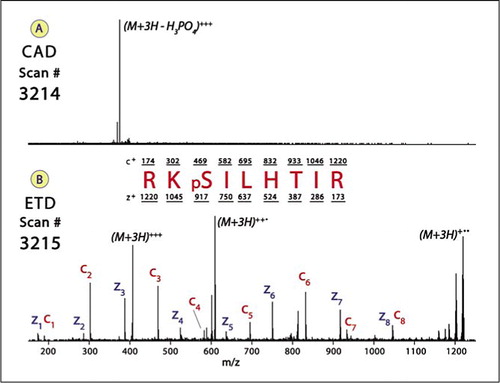
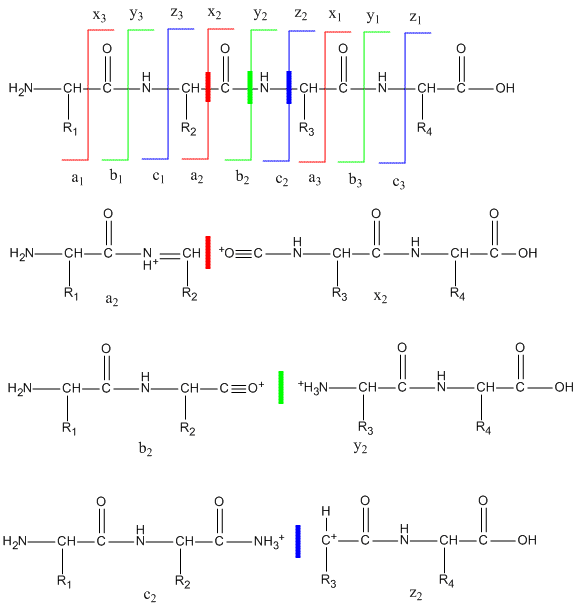
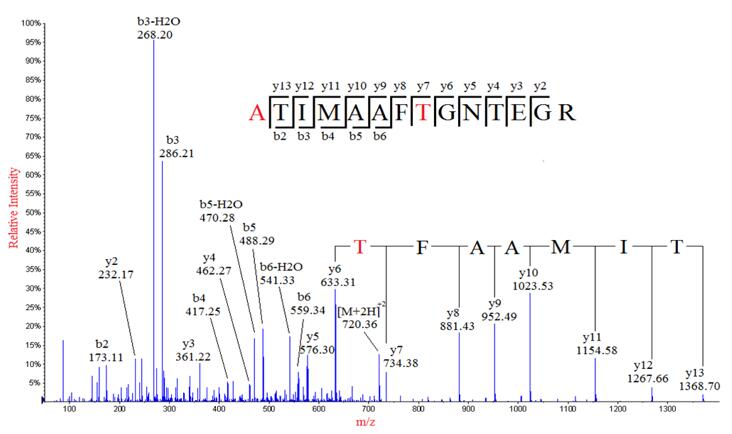
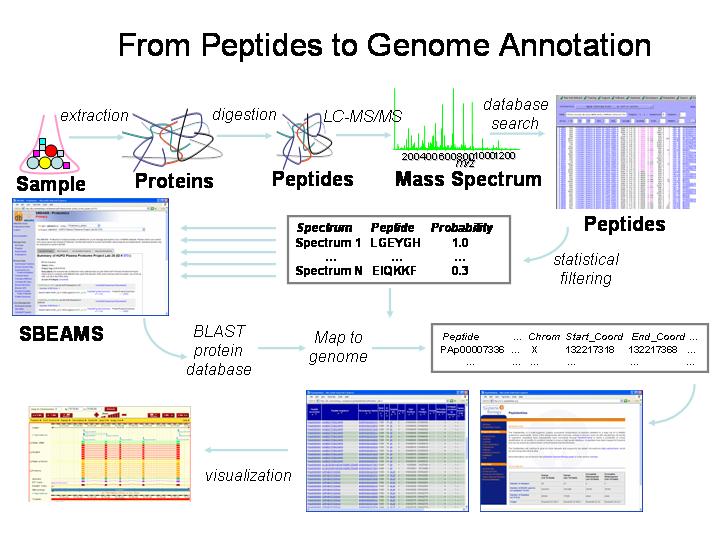
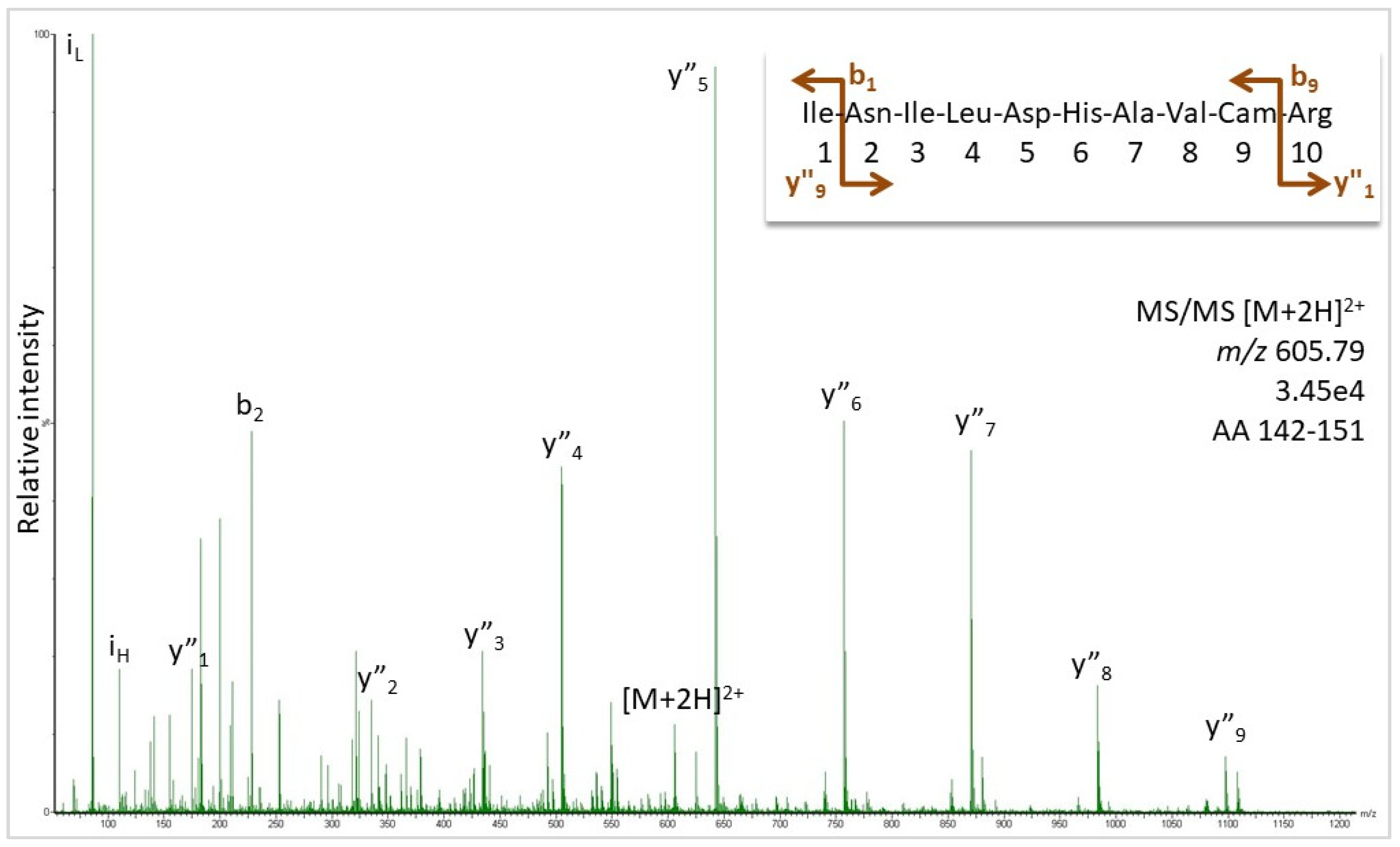
Fig. S2. Representative peptide sequencing data by MS/MS. Product ion... | Download Scientific Diagram
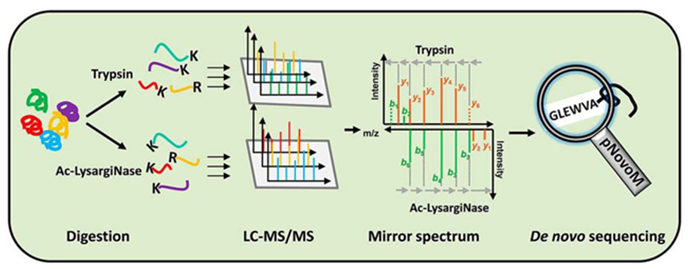
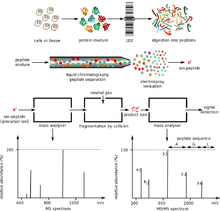
Molecules | Free Full-Text | The Current State-of-the-Art Identification of Unknown Proteins Using Mass Spectrometry Exemplified on De Novo Sequencing of a Venom Protease from Bothrops moojeni

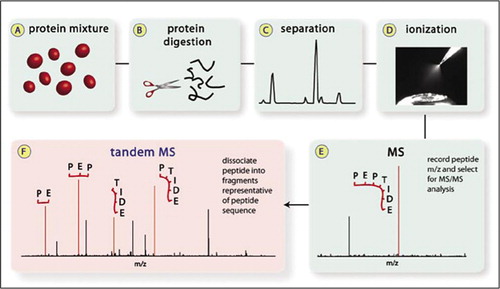
Identification and Sequencing of N-Terminal Peptides in Proteins by LC-Fluorescence-MS/MS: An Approach to Replacement of the Edman Degradation | Analytical Chemistry
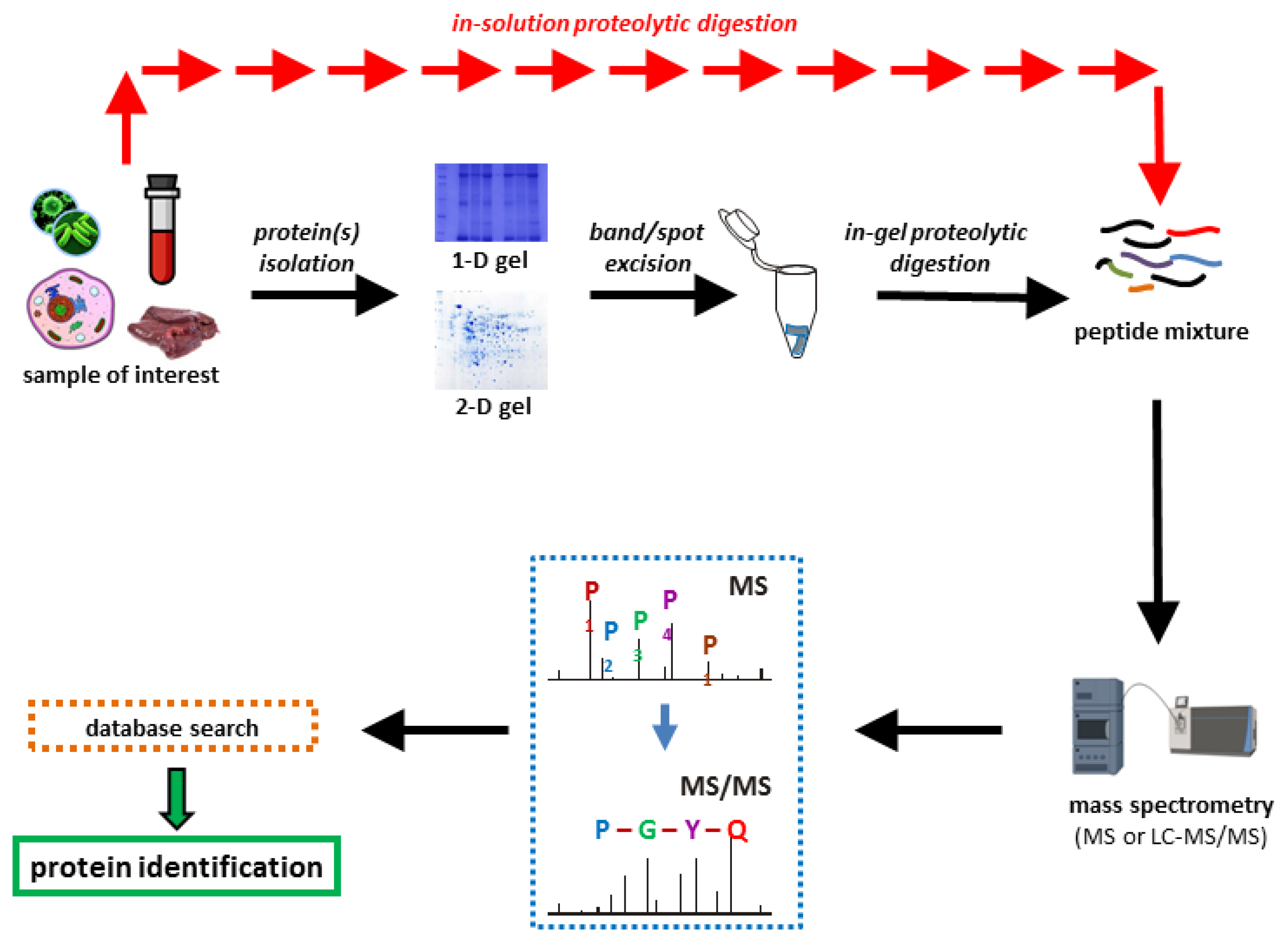
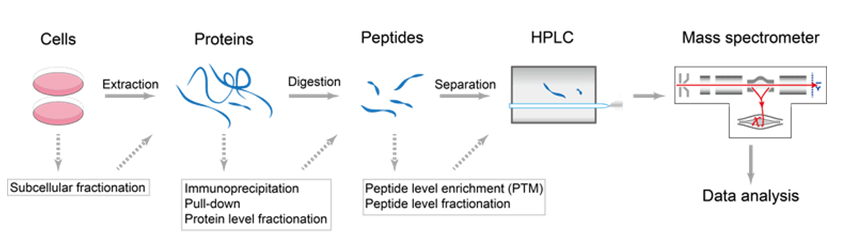
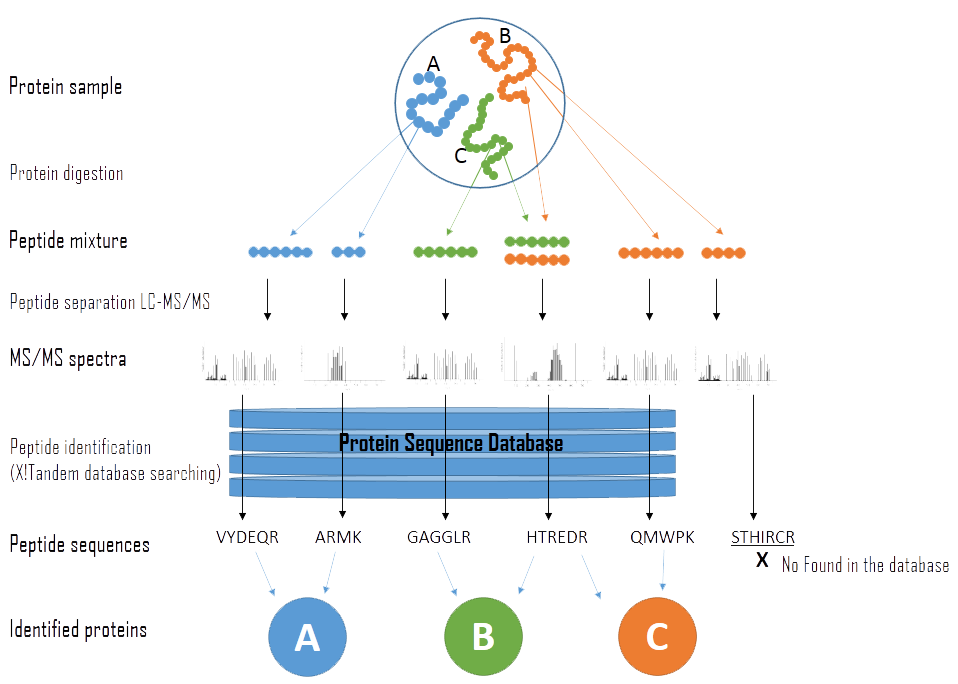
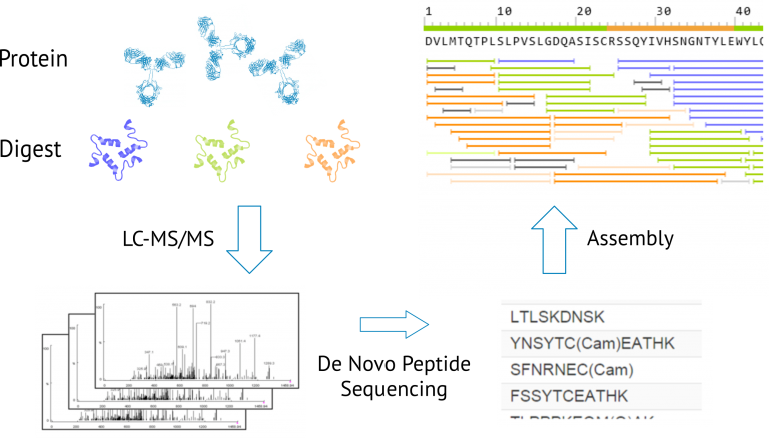
Proteomes | Free Full-Text | A Critical Review of Bottom-Up Proteomics: The Good, the Bad, and the Future of This Field
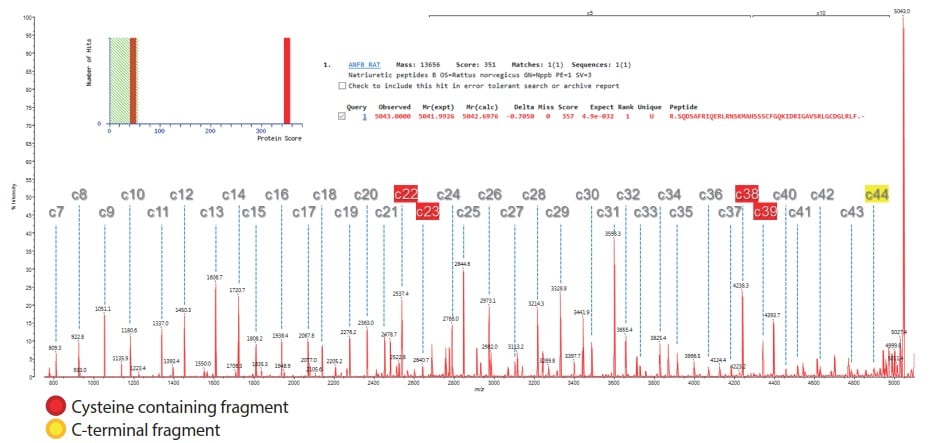
N-terminal Amino Acid Sequencing Analysis by MALDI-TOF MS/Protein Sequencer : SHIMADZU (Shimadzu Corporation)