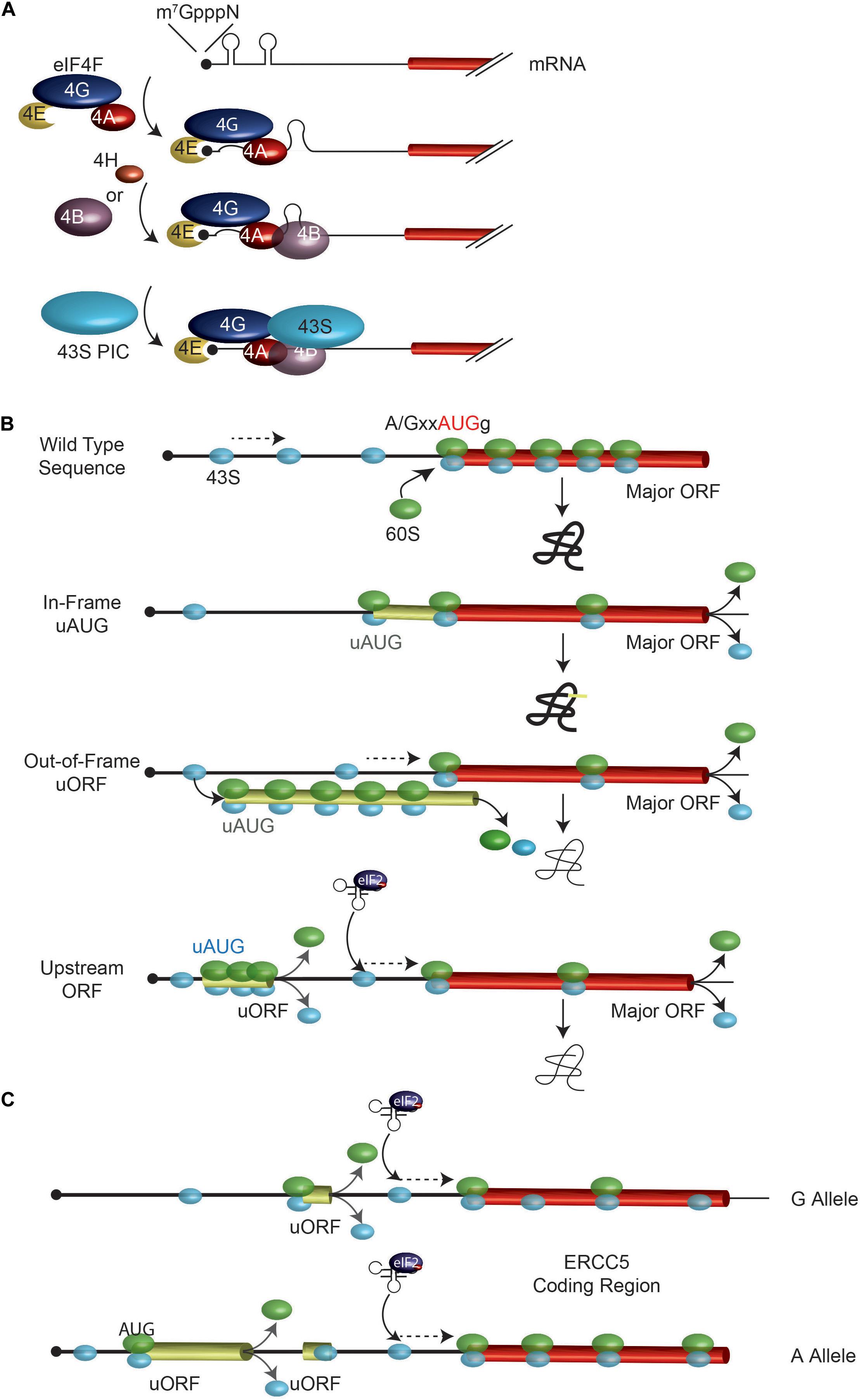
Machine learning predicts translation initiation sites in neurologic diseases with nucleotide repeat expansions | PLOS ONE

A non-coding A-to-T Kozak site change related to the transmissibility of Alpha, Delta and Omicron VOCs - Abstract - Europe PMC

Human 5' UTR design and variant effect prediction from a massively parallel translation assay. - Abstract - Europe PMC

Nucleotides upstream of the Kozak sequence strongly influence gene expression in the yeast S. cerevisiae | Journal of Biological Engineering | Full Text

IJMS | Free Full-Text | Kozak Similarity Score Algorithm Identifies Alternative Translation Initiation Codons Implicated in Cancers
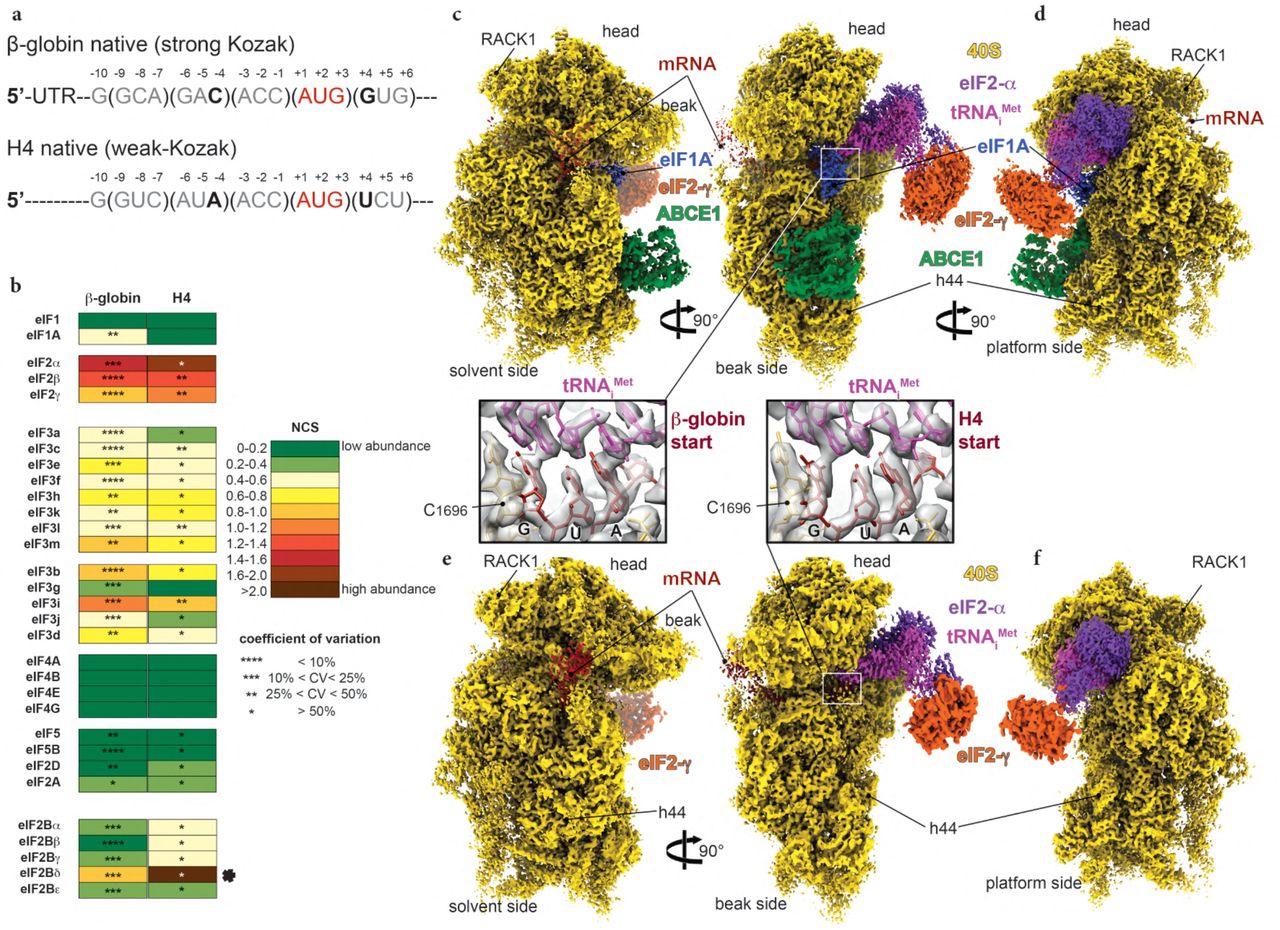
Structural insights into the Kozak sequence interactions within the mammalian late-stage initiation complexes | bioRxiv

Translation of small downstream ORFs enhances translation of canonical main open reading frames | The EMBO Journal

Analysis of Kozak consensus sequences surrounding the start codon AUG... | Download Scientific Diagram

Fine-tuning the expression of pathway gene in yeast using a regulatory library formed by fusing a synthetic minimal promoter with different Kozak variants | Microbial Cell Factories | Full Text
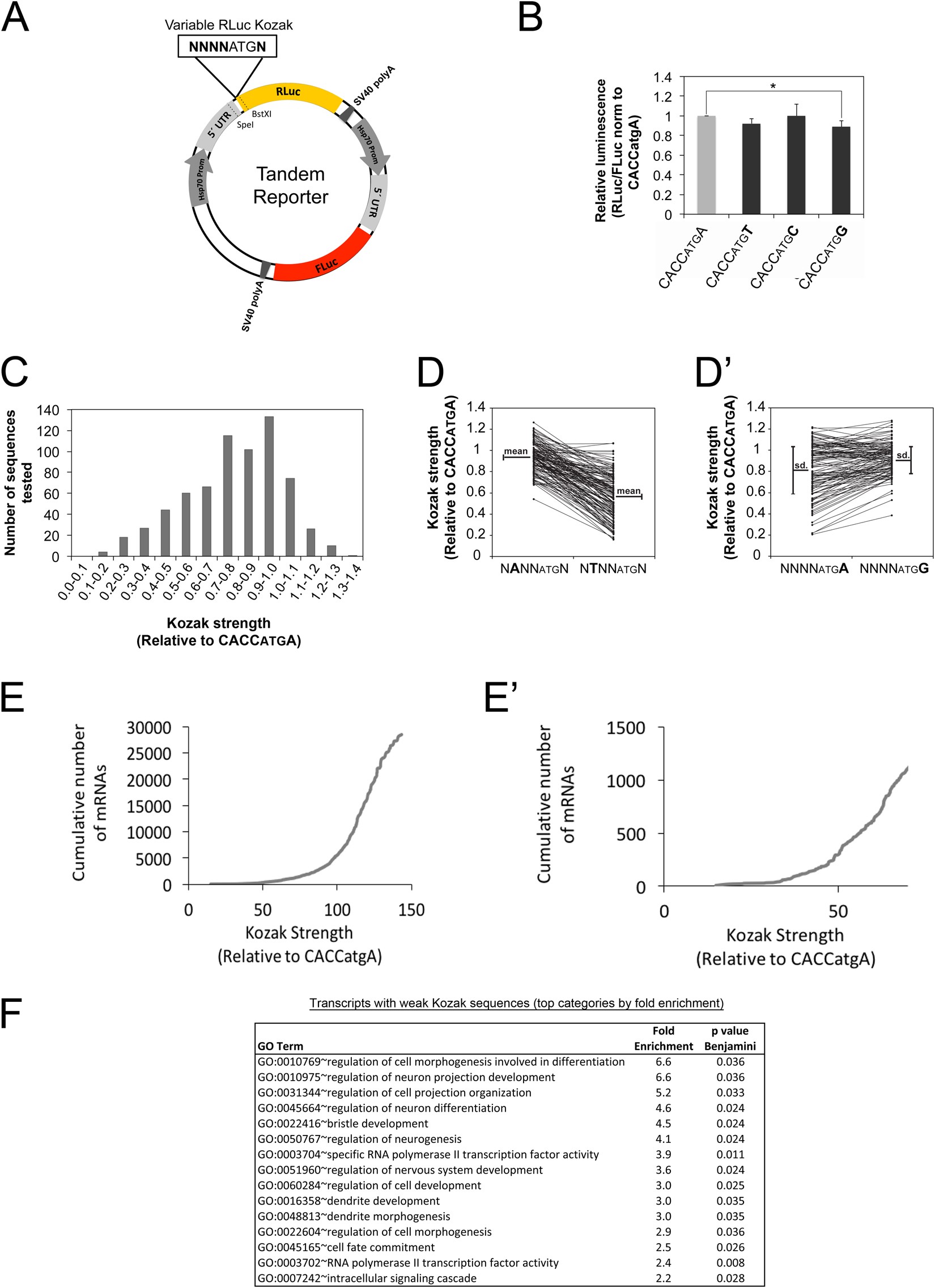
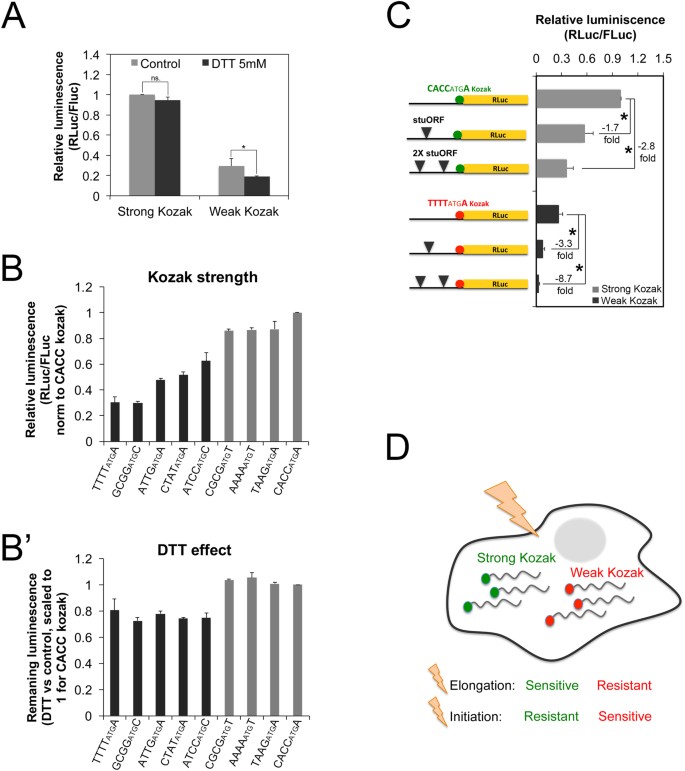
Changes in global translation elongation or initiation rates shape the proteome via the Kozak sequence | Scientific Reports

An expanded toolkit for Drosophila gene tagging using synthesized homology donor constructs for CRISPR mediated homologous recombination | bioRxiv

Relationship between consensus Kozak sequence at the true start ATG for... | Download Scientific Diagram