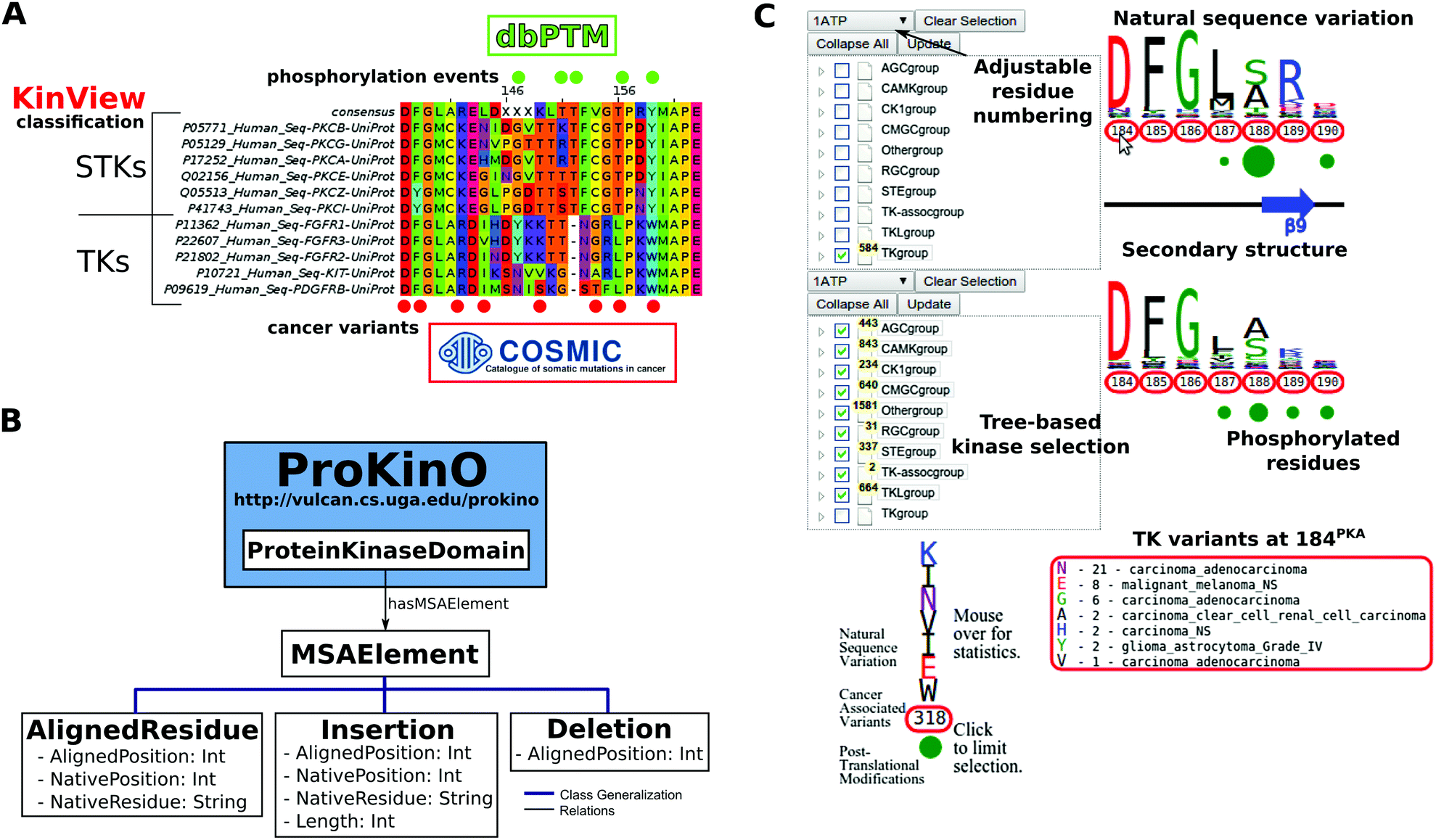
KinView: a visual comparative sequence analysis tool for integrated kinome research - Molecular BioSystems (RSC Publishing) DOI:10.1039/C6MB00466K

Life | Free Full-Text | Structural Insights into Protein Regulation by Phosphorylation and Substrate Recognition of Protein Kinases/Phosphatases
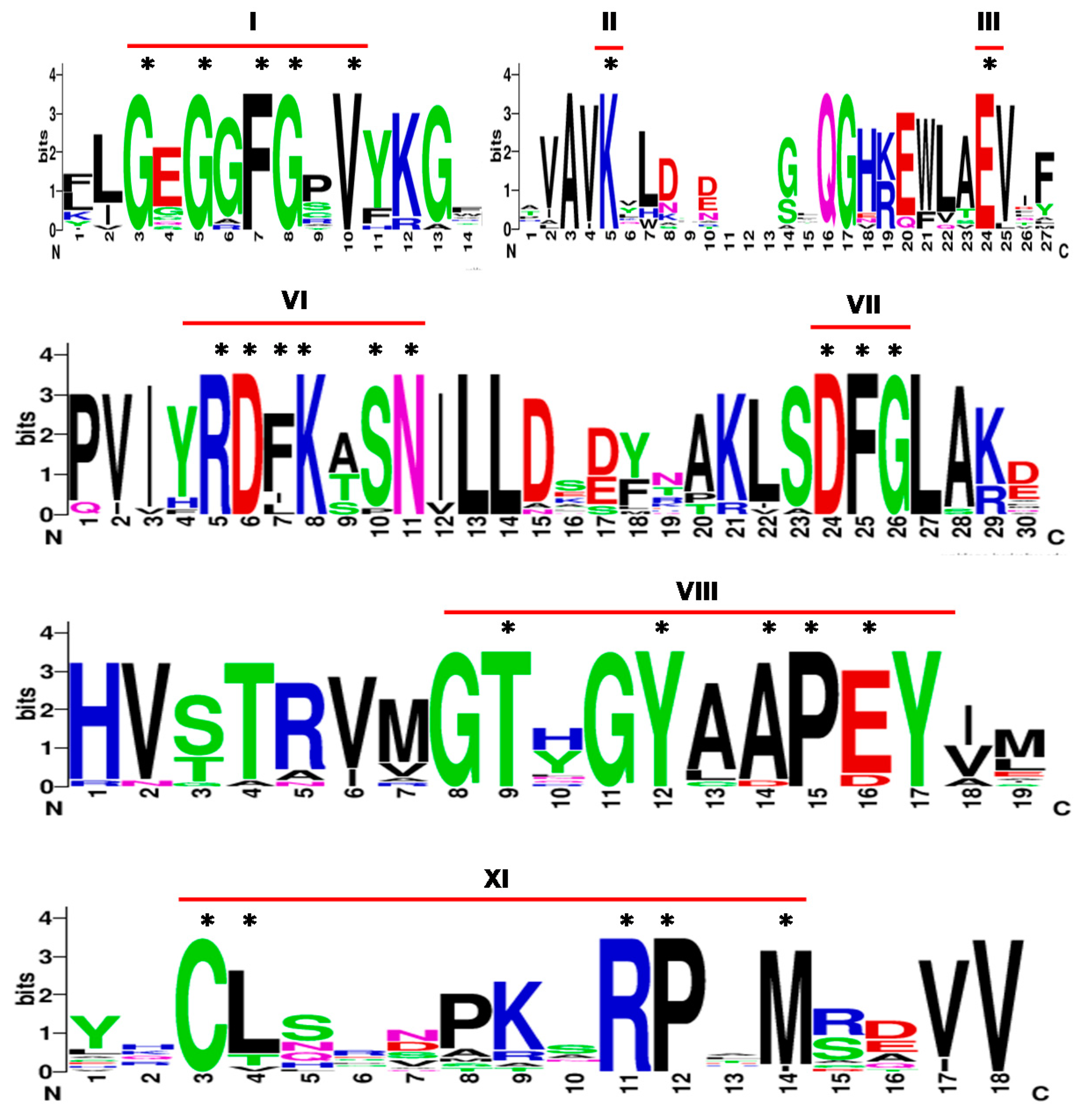
Plants | Free Full-Text | Characterization of Atypical Protein Tyrosine Kinase (PTK) Genes and Their Role in Abiotic Stress Response in Rice

KinasePhos 3.0: Redesign and Expansion of the Prediction on Kinase-specific Phosphorylation Sites | bioRxiv

Determination of a consensus motif for Plk1 phosphorylation . A–C, the... | Download Scientific Diagram

The consensus sequences of Protein kinase domain (Pkinase, pfam00069)... | Download Scientific Diagram
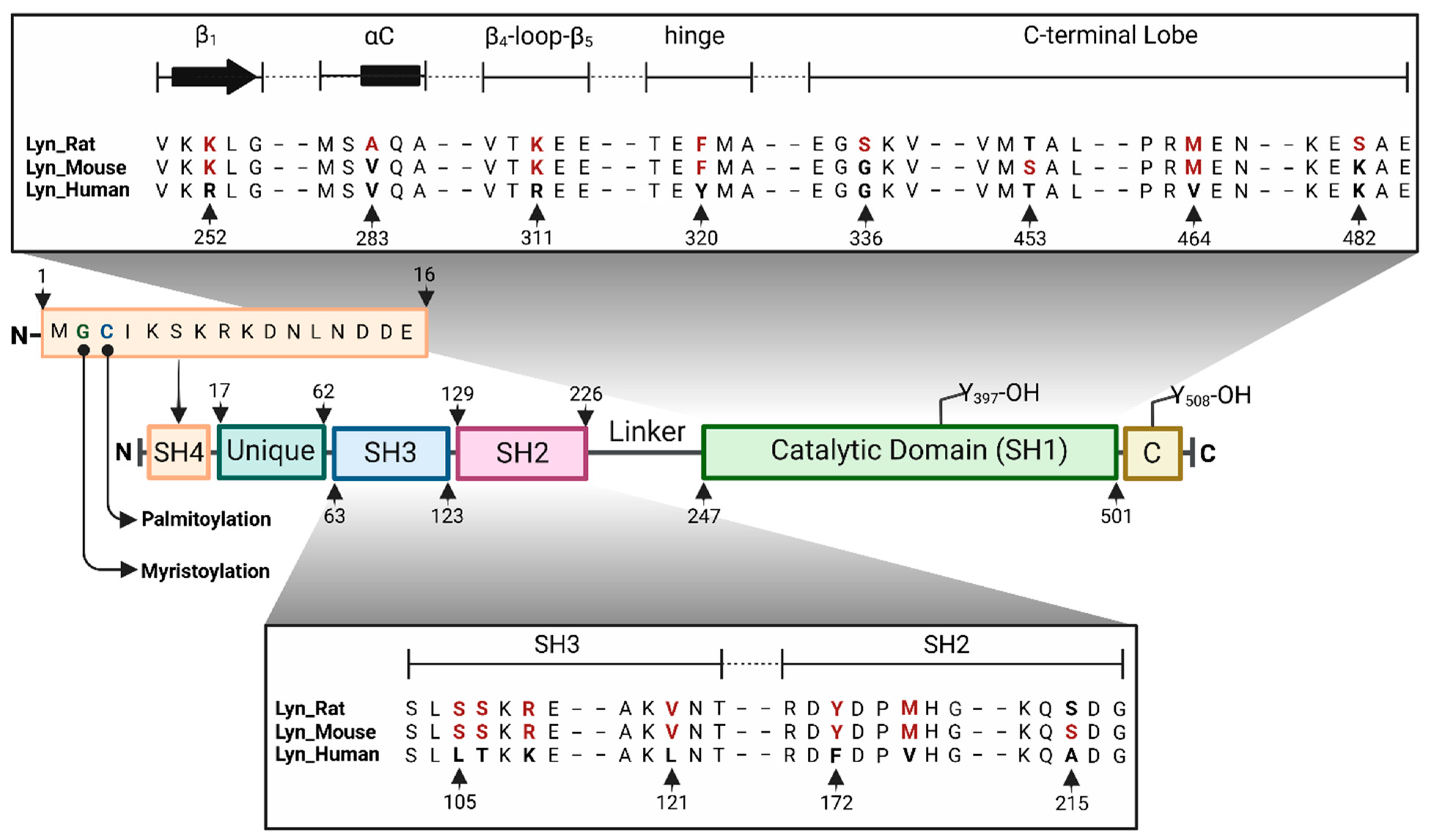
Kinases and Phosphatases | Free Full-Text | Lyn Kinase Structure, Regulation, and Involvement in Neurodegenerative Diseases: A Mini Review

Kinase consensus sequences. Consensus sequences of some of the kinases... | Download Scientific Diagram
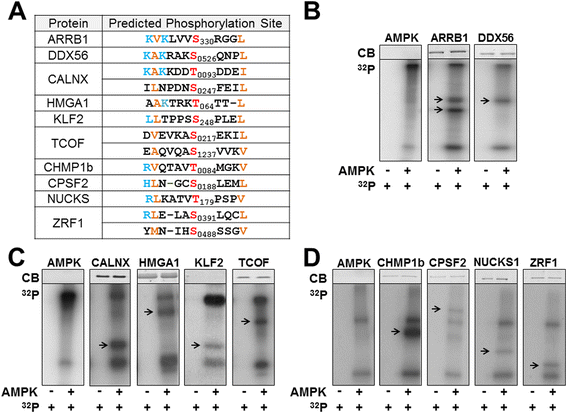
Identification of AMP-activated protein kinase targets by a consensus sequence search of the proteome | BMC Systems Biology | Full Text

Kinase consensus sequences. Consensus sequences of some of the kinases... | Download Scientific Diagram

Identification of Kinases and Interactors of p53 Using Kinase-Catalyzed Cross-Linking and Immunoprecipitation | Journal of the American Chemical Society

An illustration of extraction of different consensus sequence motifs by... | Download Scientific Diagram

Substrate Recognition Mechanism of Atypical Protein Kinase Cs Revealed by the Structure of PKCι in Complex with a Substrate Peptide from Par-3 - ScienceDirect
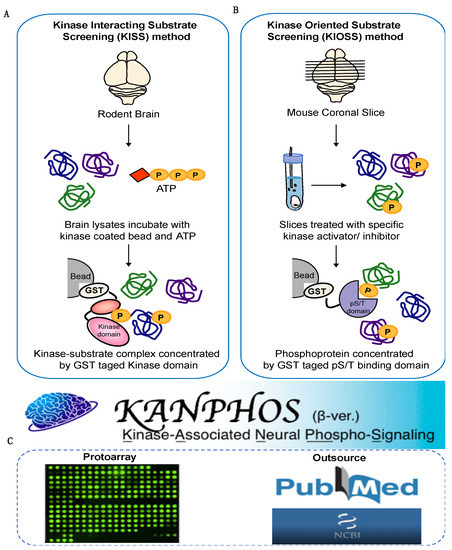
Cells | Free Full-Text | KANPHOS: A Database of Kinase-Associated Neural Protein Phosphorylation in the Brain

A Structurally-Validated Multiple Sequence Alignment of 497 Human Protein Kinase Domains | Scientific Reports
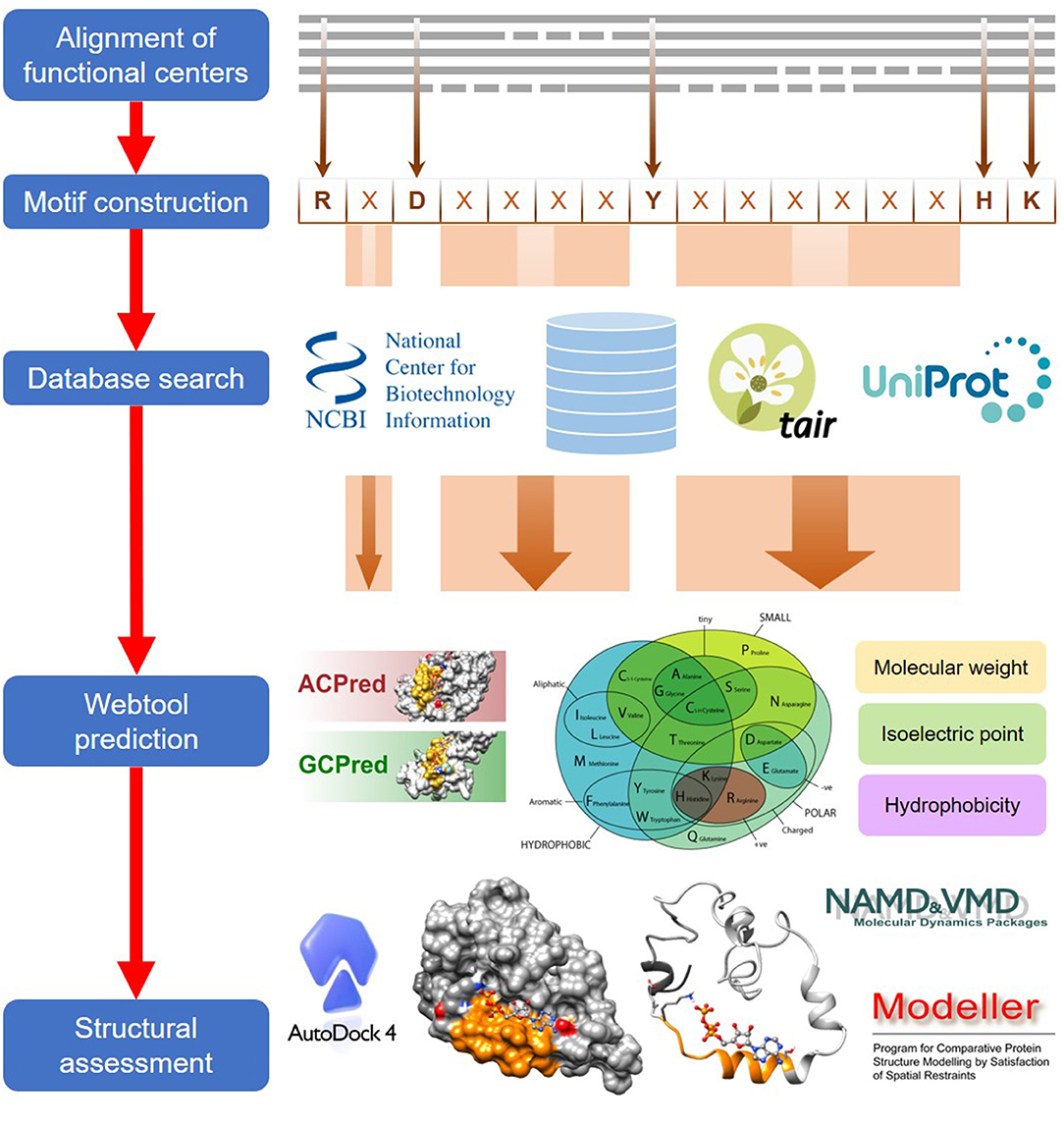
Frontiers | Computational Identification of Functional Centers in Complex Proteins: A Step-by-Step Guide With Examples
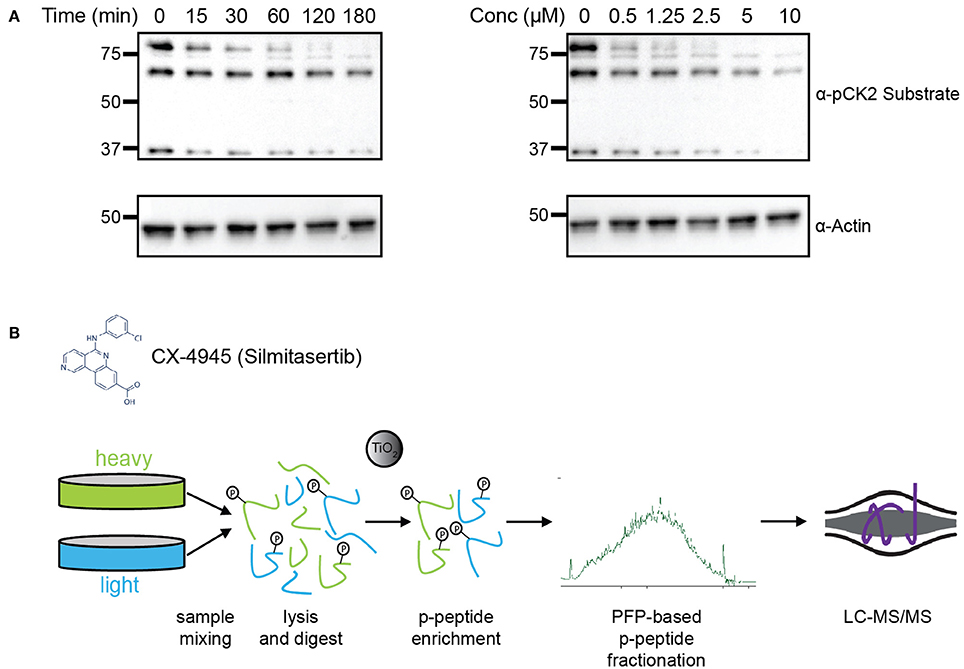
Frontiers | Identification of Candidate Casein Kinase 2 Substrates in Mitosis by Quantitative Phosphoproteomics