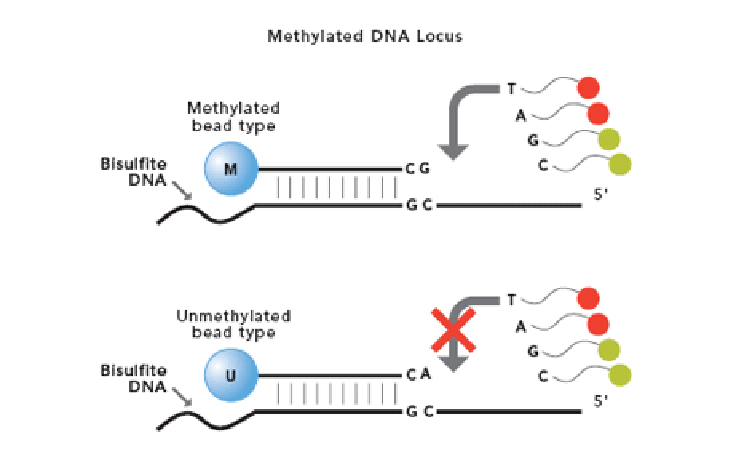
High-resolution targeted bisulfite sequencing reveals blood cell type-specific DNA methylation patterns in IL13 and ORMDL3 | Clinical Epigenetics | Full Text
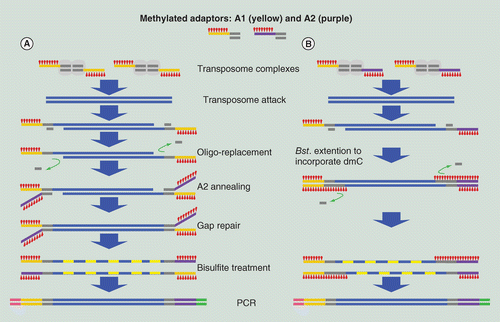
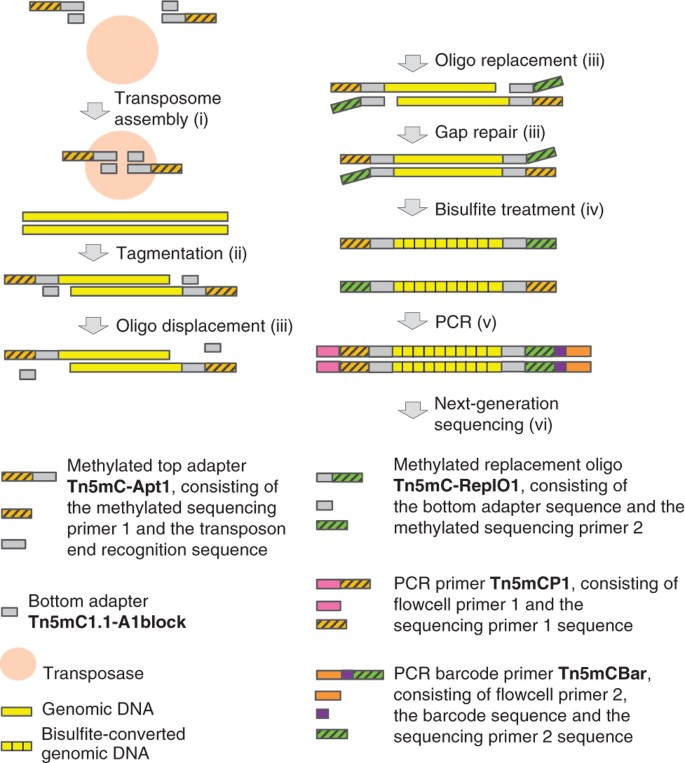
Improved tagmentation-based whole-genome bisulfite sequencing for input DNA from less than 100 mammalian cells | Epigenomics

Single-Cell DNA Methylome Sequencing and Bioinformatic Inference of Epigenomic Cell-State Dynamics - ScienceDirect
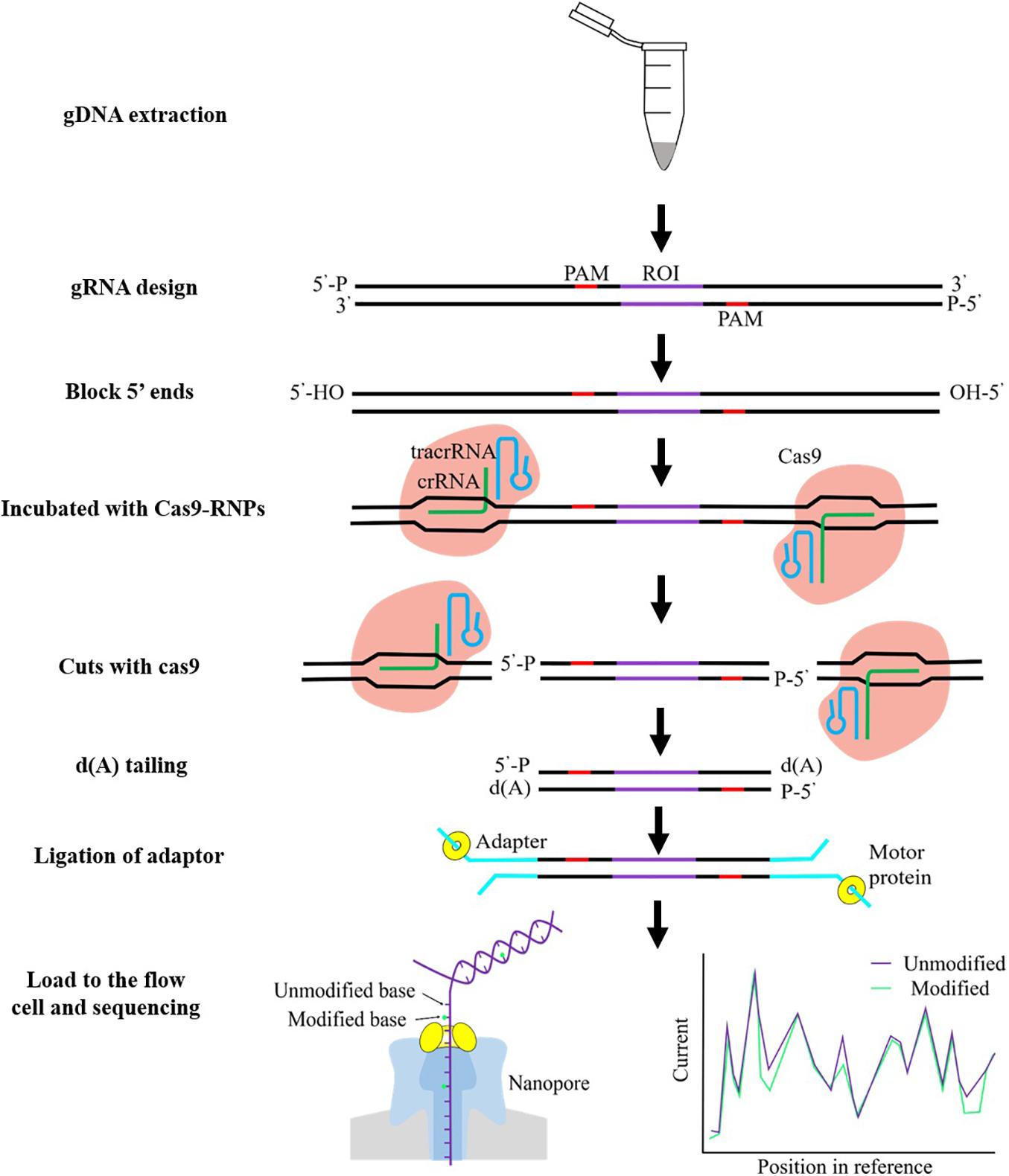
Frontiers | Cancer Biomarkers Discovery of Methylation Modification With Direct High-Throughput Nanopore Sequencing
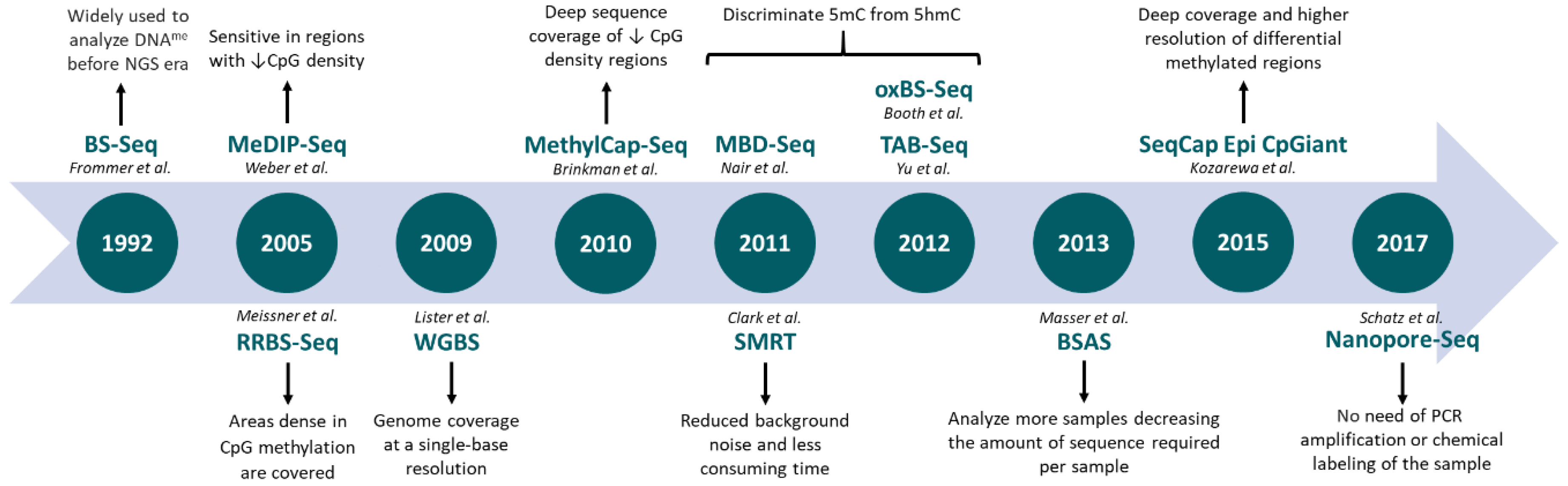
Genes | Free Full-Text | Profiling DNA Methylation Based on Next-Generation Sequencing Approaches: New Insights and Clinical Applications
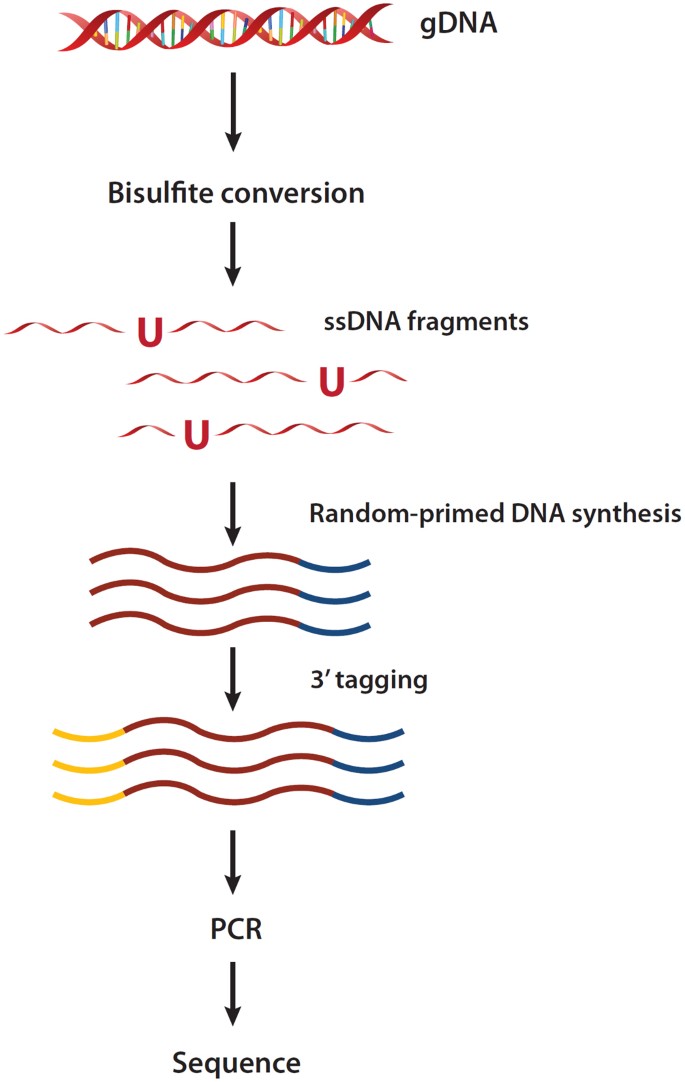
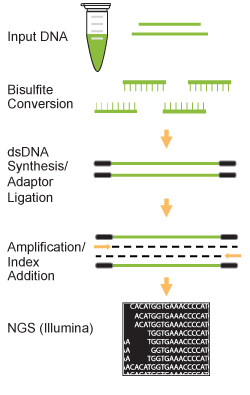
EpiGnome™ Methyl-Seq Kit: a novel post–bisulfite conversion library prep method for methylation analysis | Nature Methods

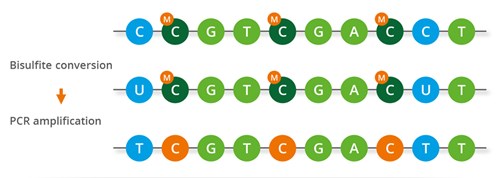
Brush Up: What Is Bisulfite Sequencing and How Do Researchers Use It to Study DNA Methylation? | The Scientist Magazine®