![PDF] Will long-read sequencing technologies replace short-read sequencing technologies in the next 10 years? | Semantic Scholar PDF] Will long-read sequencing technologies replace short-read sequencing technologies in the next 10 years? | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/1c32c3437a078ea1285ca9e19a0dcdcdf6434bc5/2-Figure1-1.png)
PDF] Will long-read sequencing technologies replace short-read sequencing technologies in the next 10 years? | Semantic Scholar

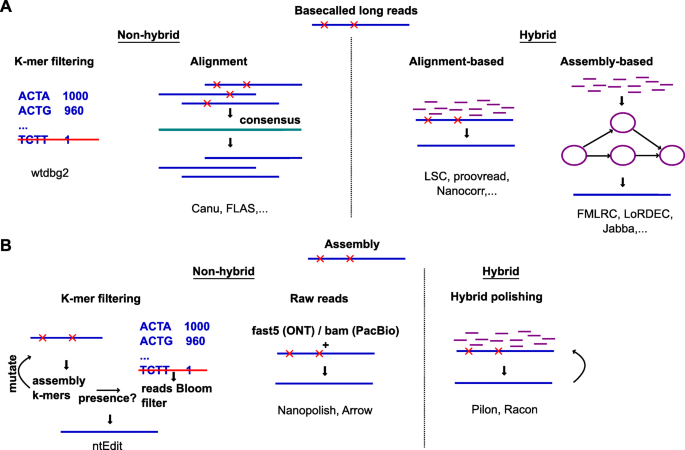
Tools and Strategies for Long-Read Sequencing and De Novo Assembly of Plant Genomes: Trends in Plant Science

Multiplexed Assembly and Annotation of Synthetic Biology Constructs Using Long-Read Nanopore Sequencing | ACS Synthetic Biology
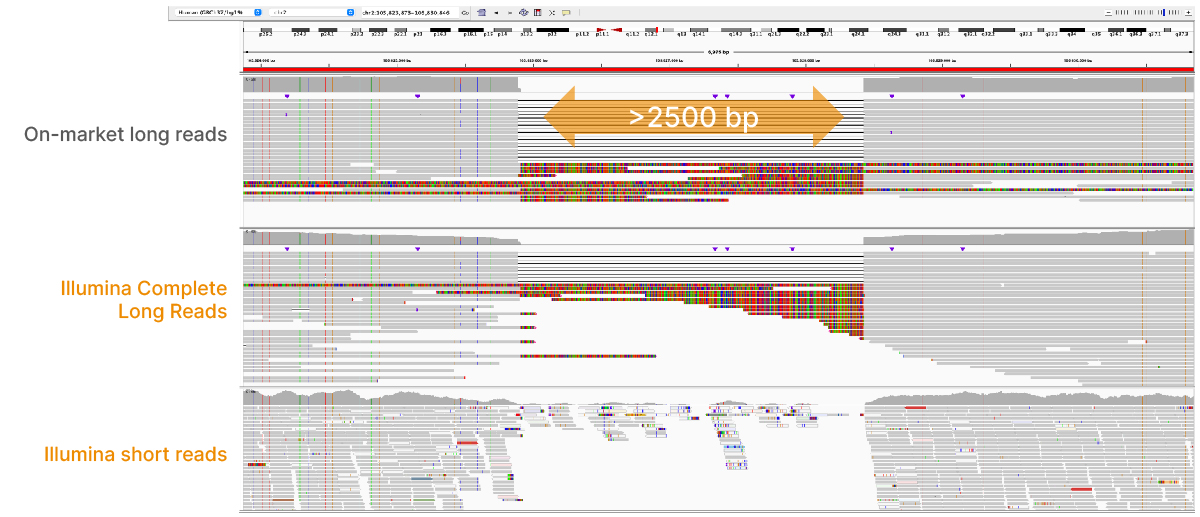
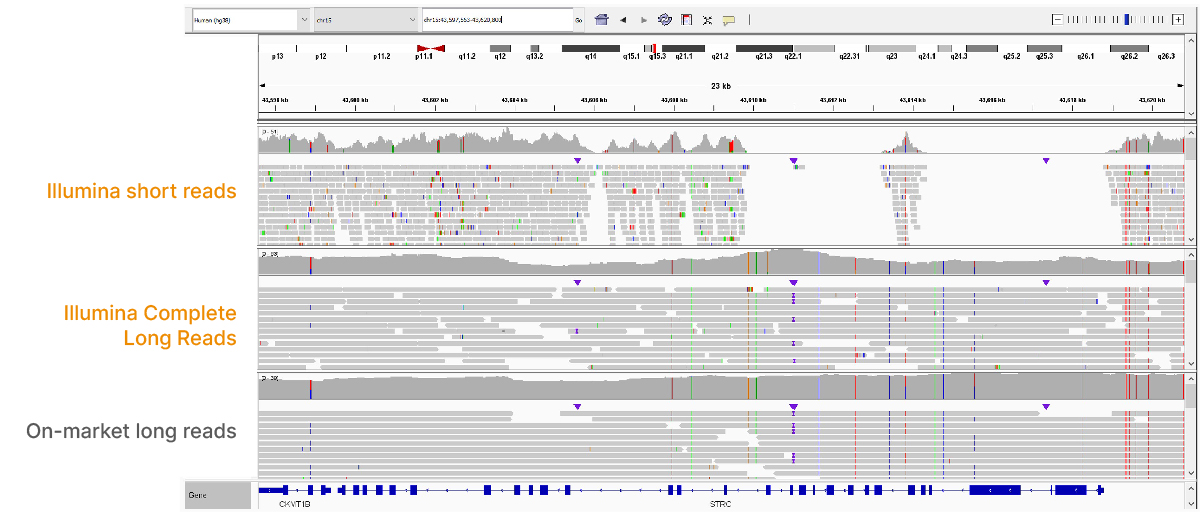
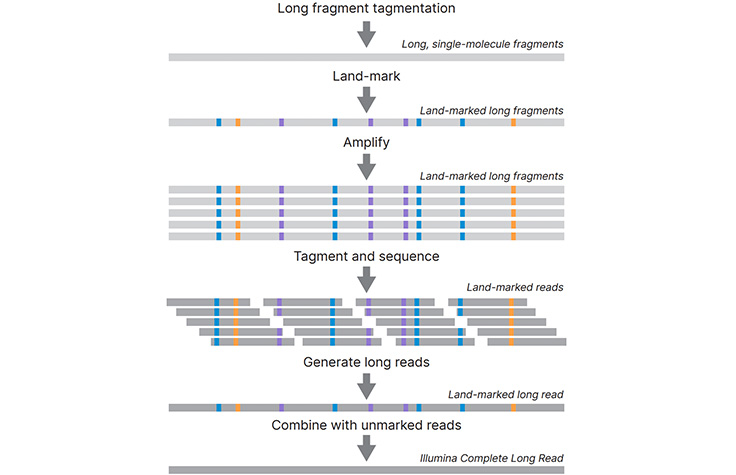
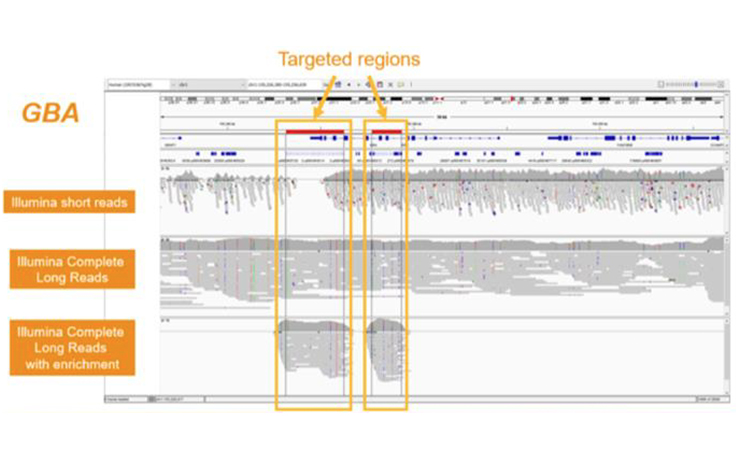
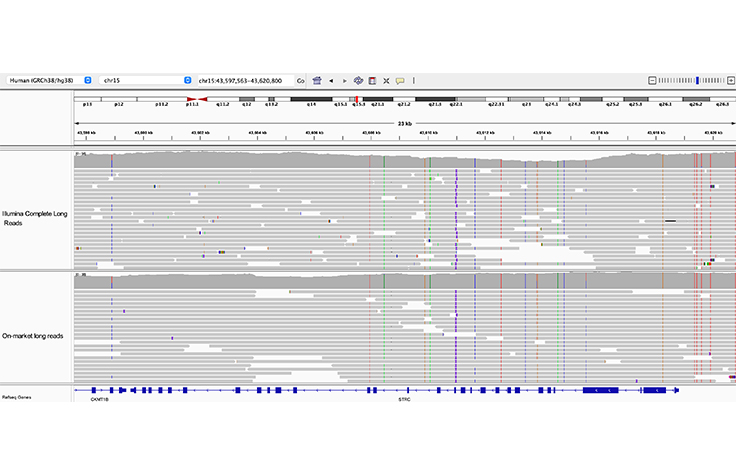
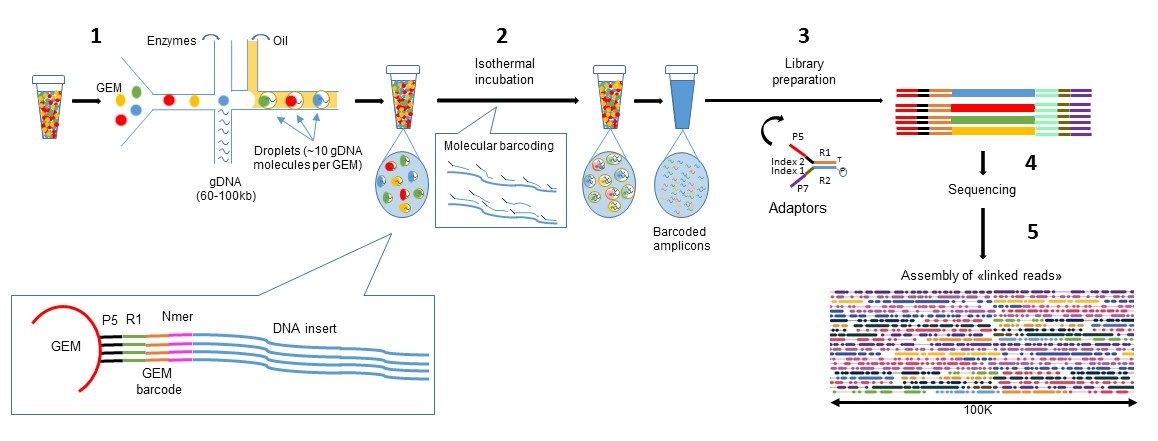
High-performance long-read assay enables contiguous data with N50 of 6–7 kb on existing Illumina platforms
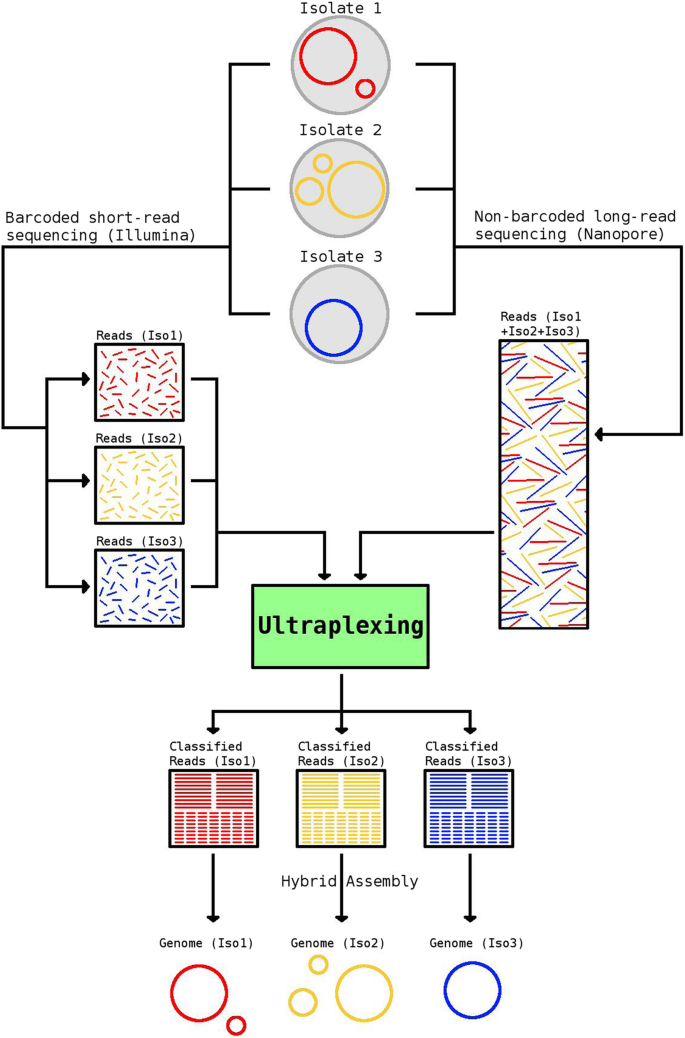
Ultraplexing: increasing the efficiency of long-read sequencing for hybrid assembly with k-mer-based multiplexing | Genome Biology | Full Text
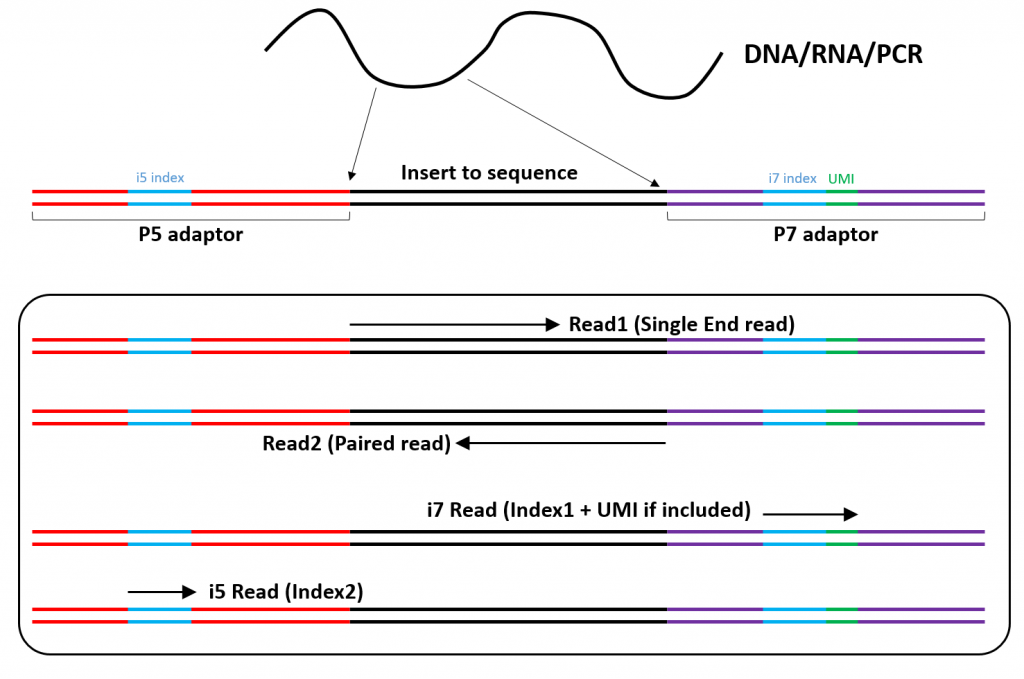
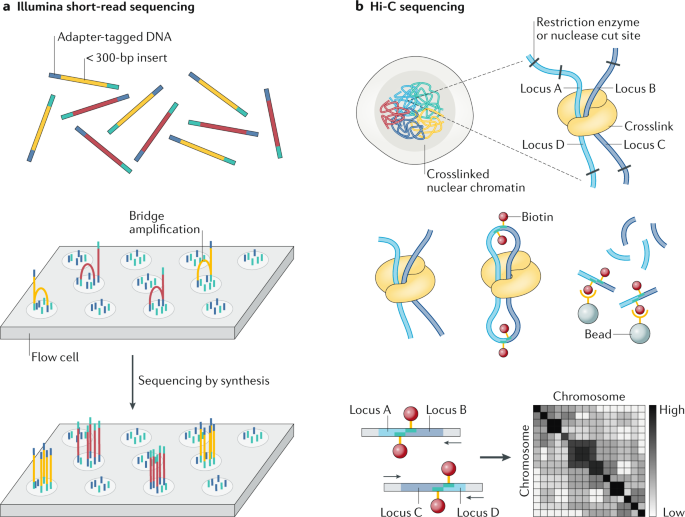
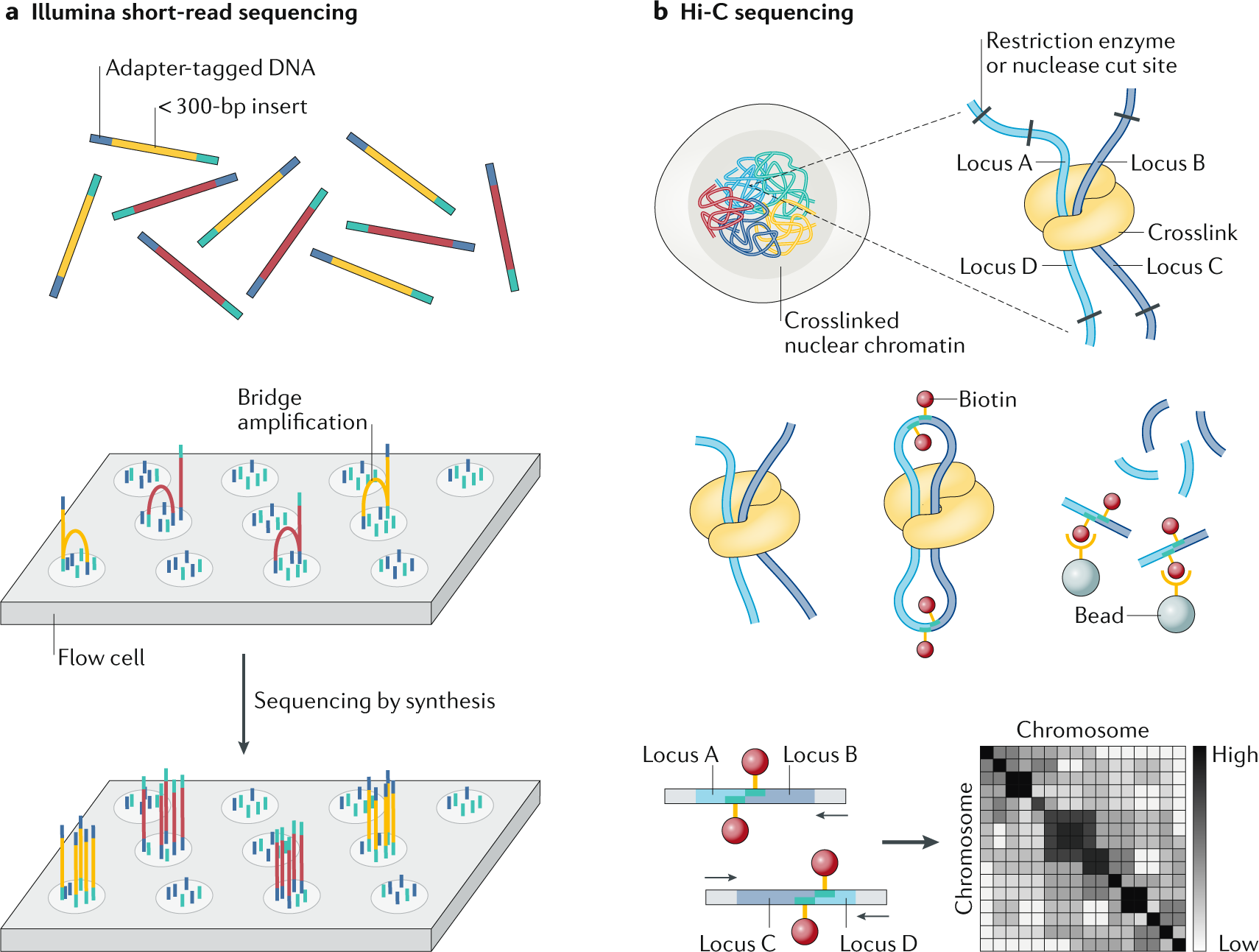
Major short-read and long-read sequencing technologies. (A) Illumina... | Download Scientific Diagram
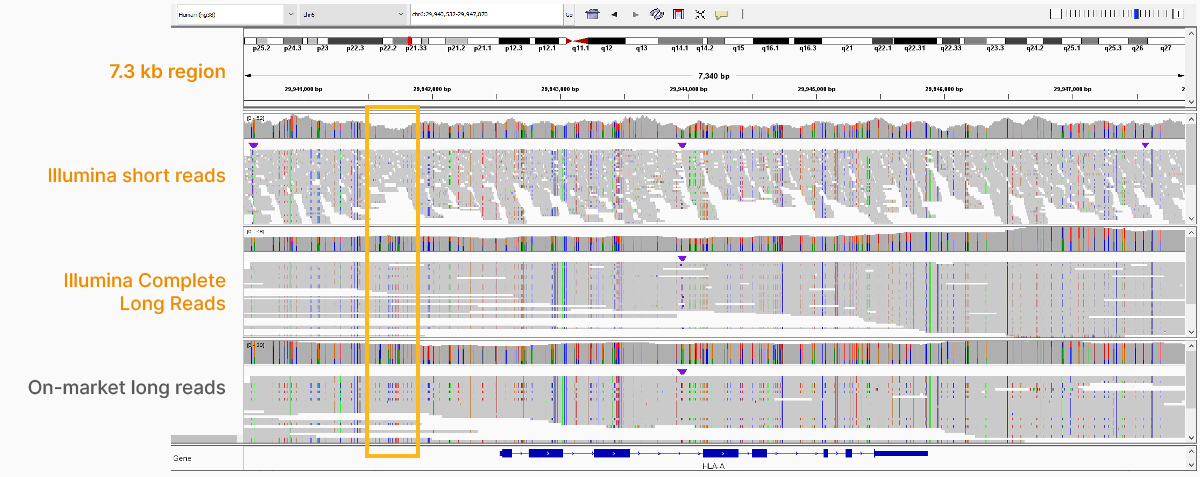
High-performance long-read assay enables contiguous data with N50 of 6–7 kb on existing Illumina platforms