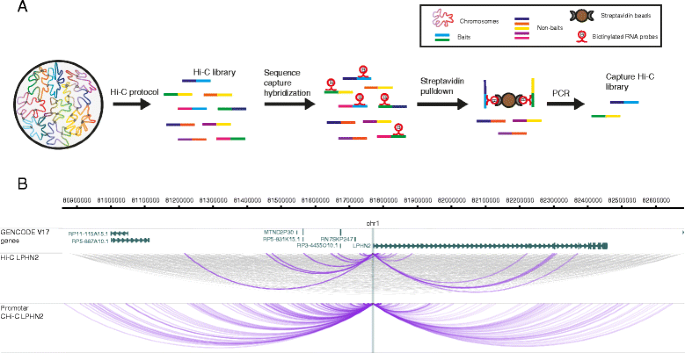
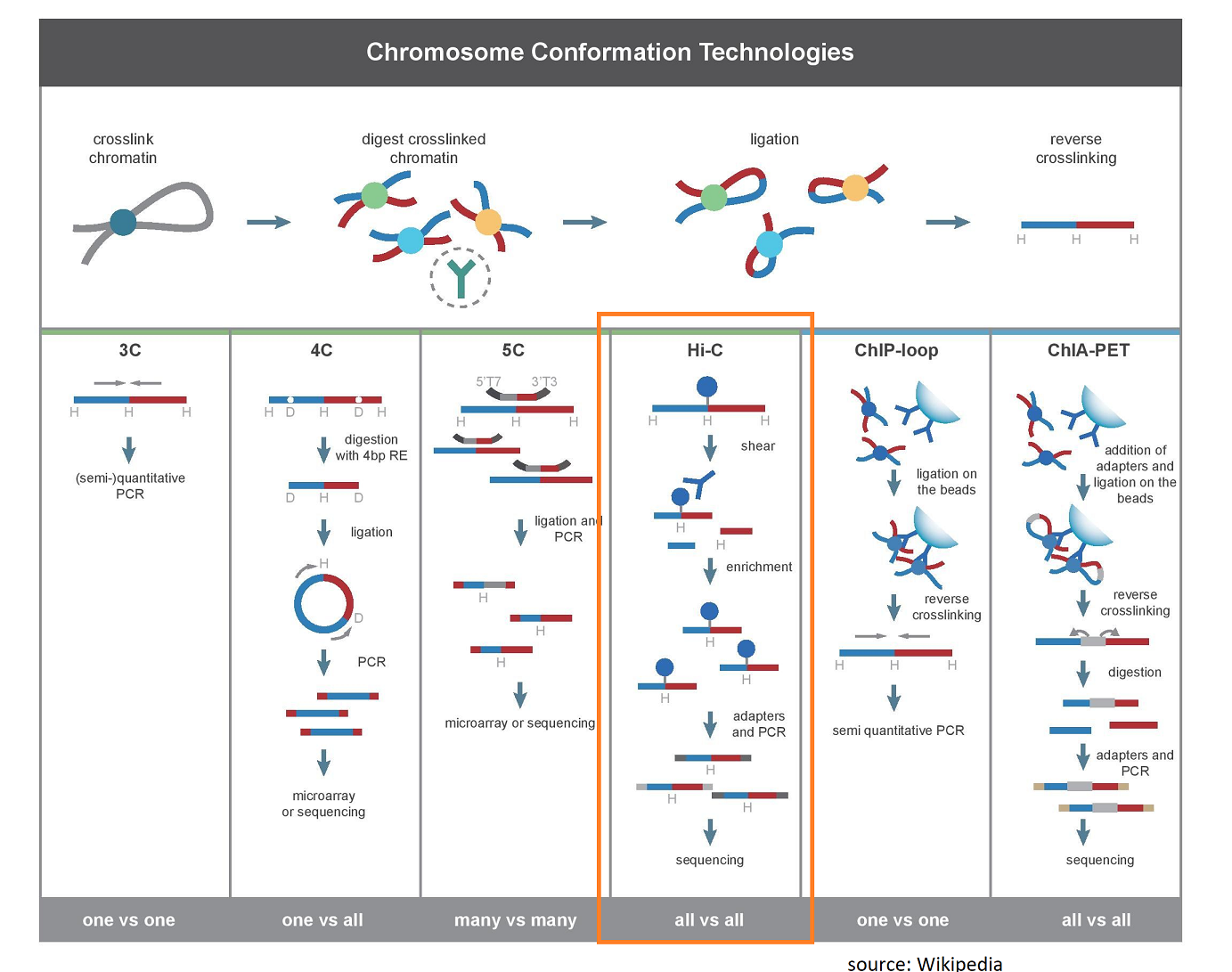
BL-Hi-C is an efficient and sensitive approach for capturing structural and regulatory chromatin interactions | Nature Communications
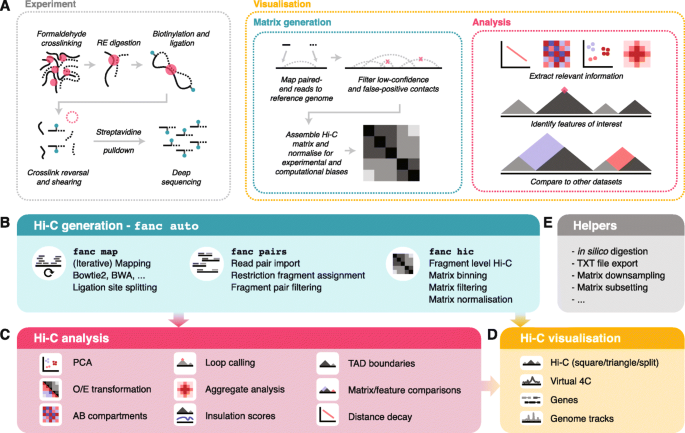
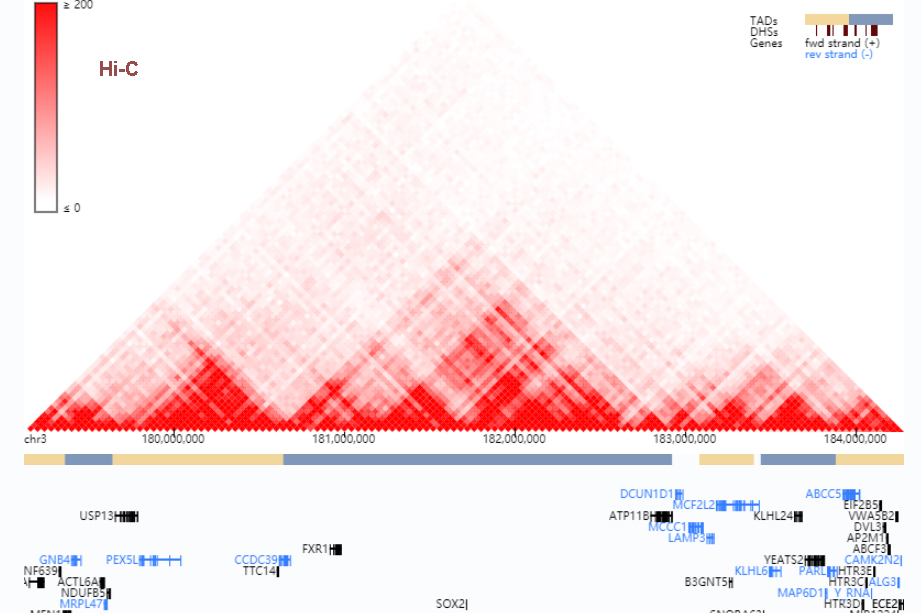
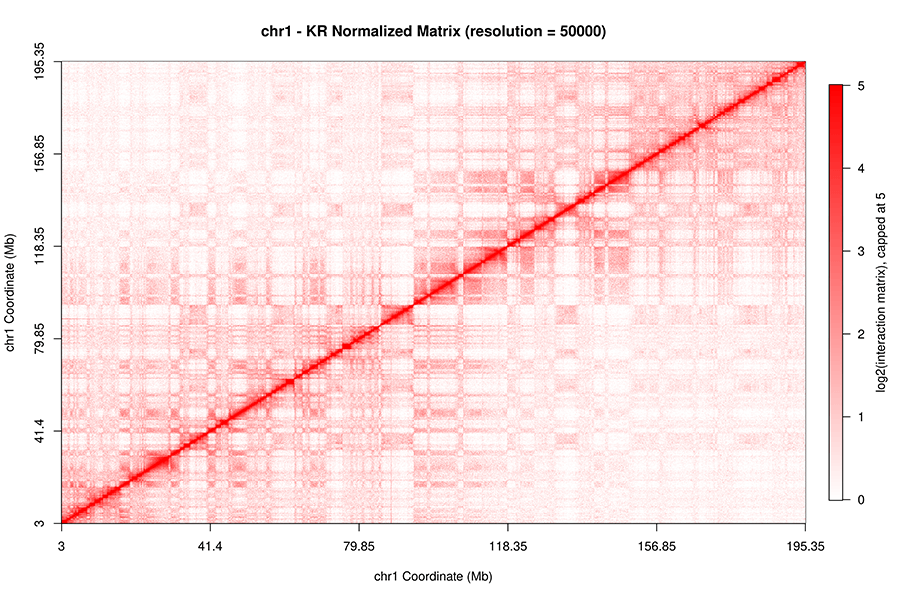
FAN-C: a feature-rich framework for the analysis and visualisation of chromosome conformation capture data | Genome Biology | Full Text

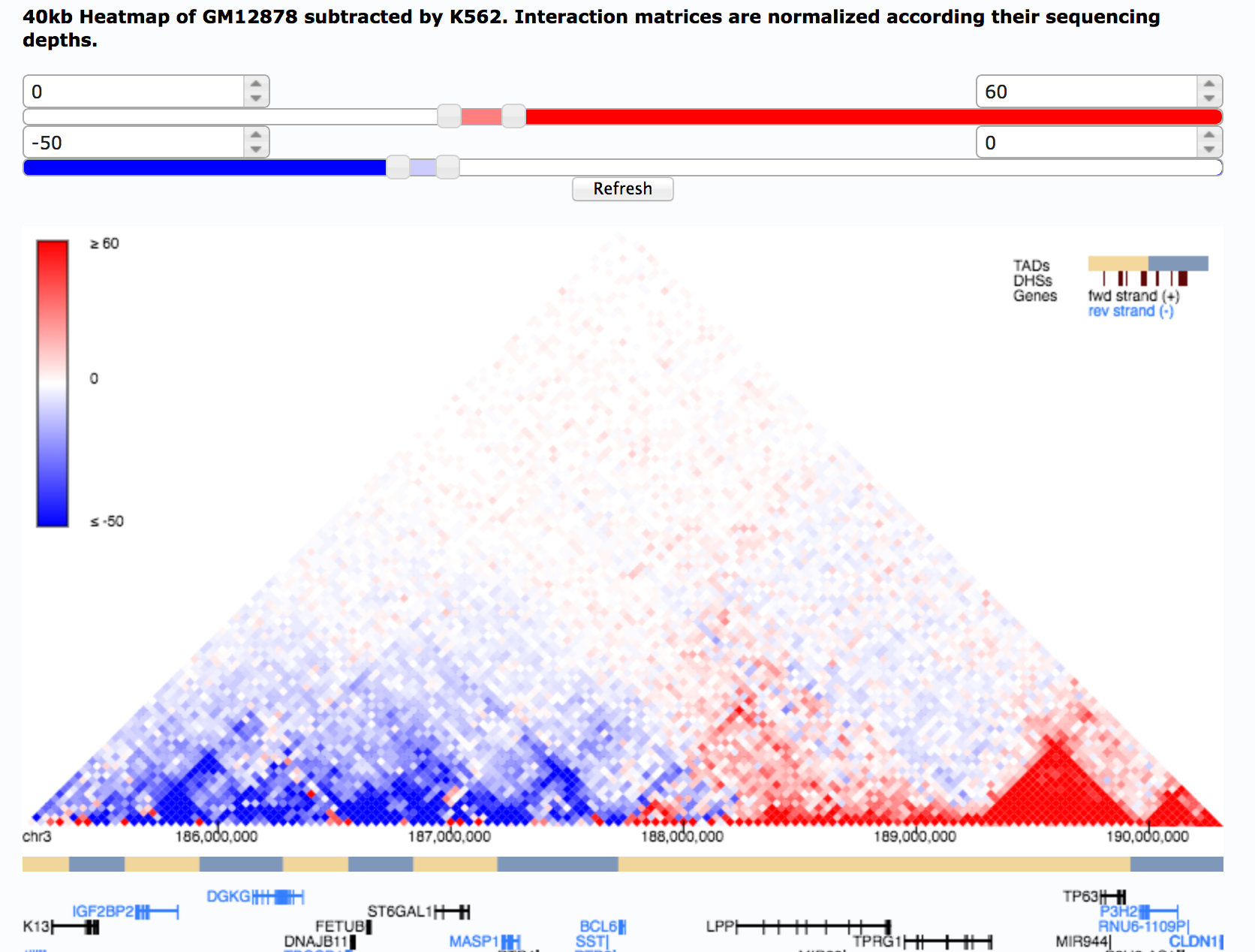
Liquid chromatin Hi-C characterizes compartment-dependent chromatin interaction dynamics | Nature Genetics

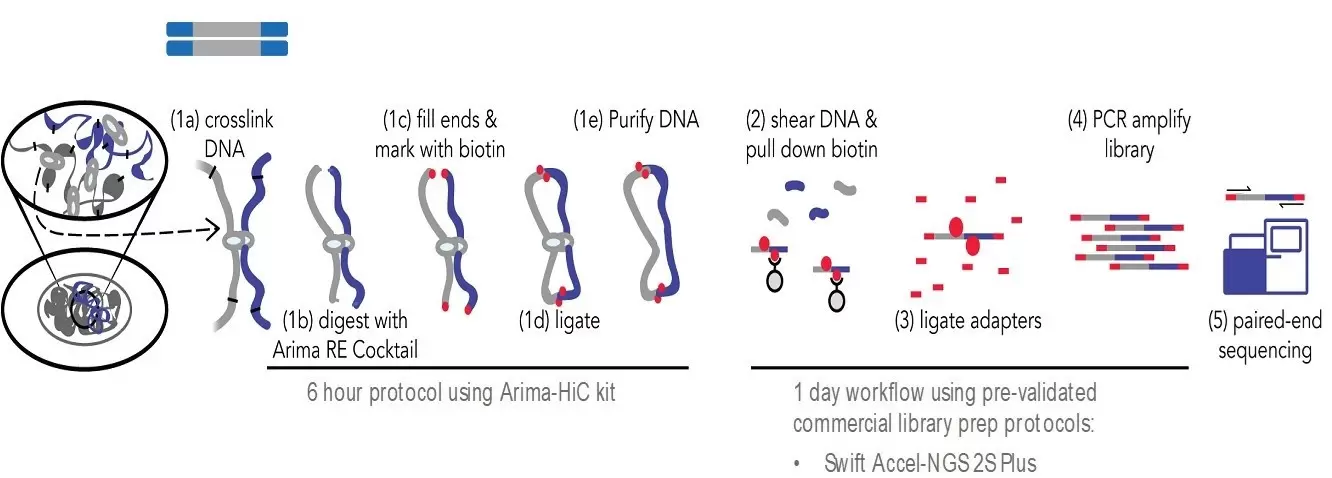
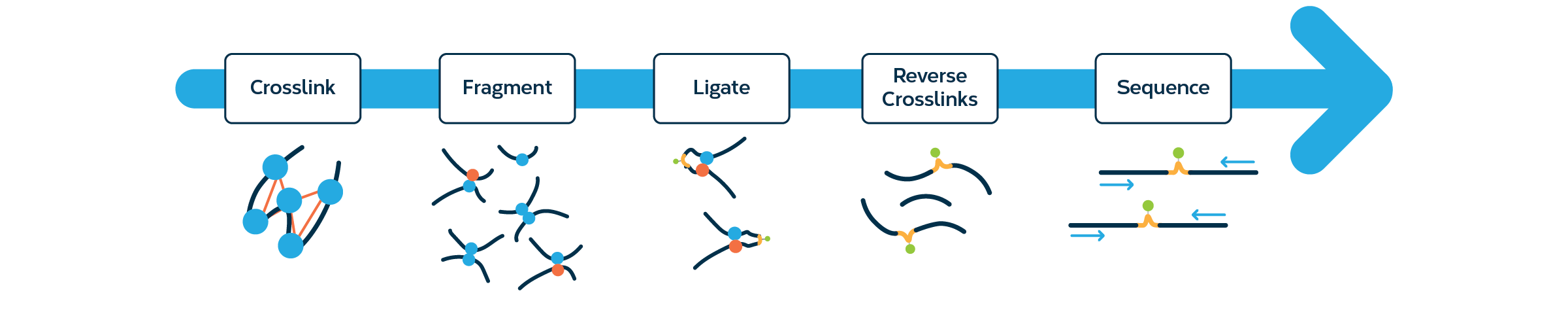
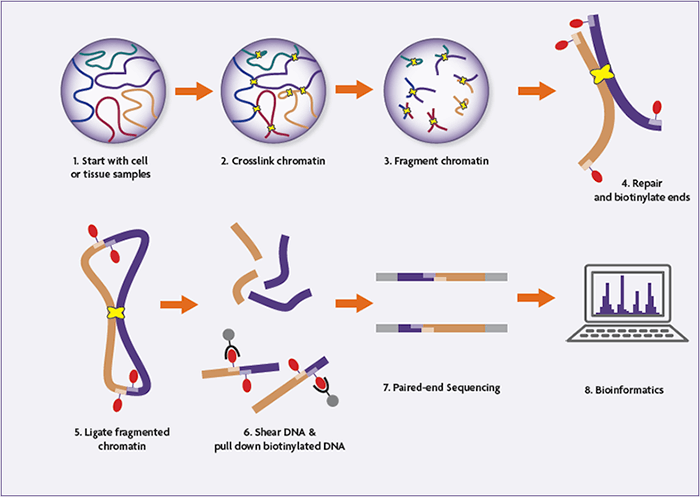
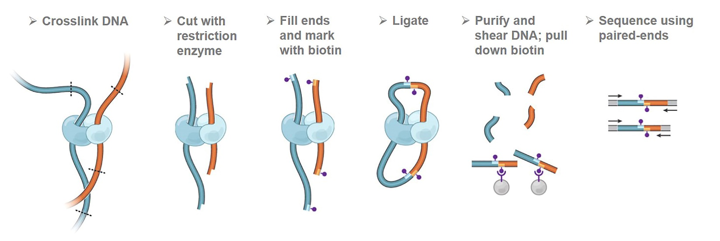
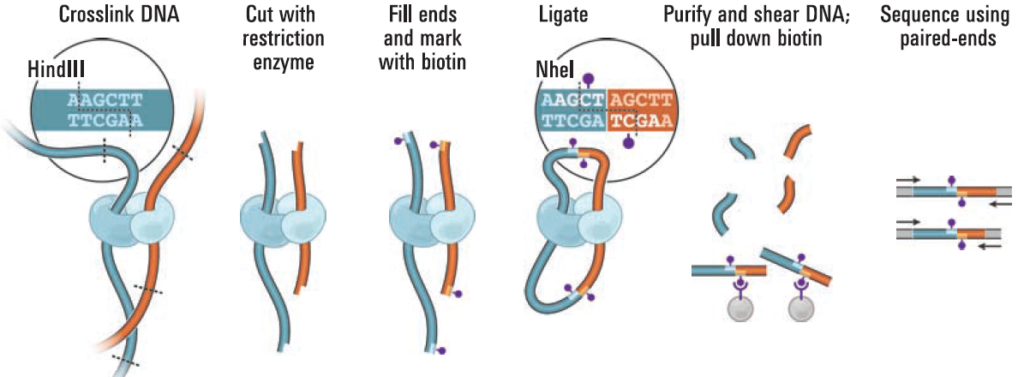
HiC workflow. Top row: Experimental steps to prepare preliminary HiC... | Download Scientific Diagram
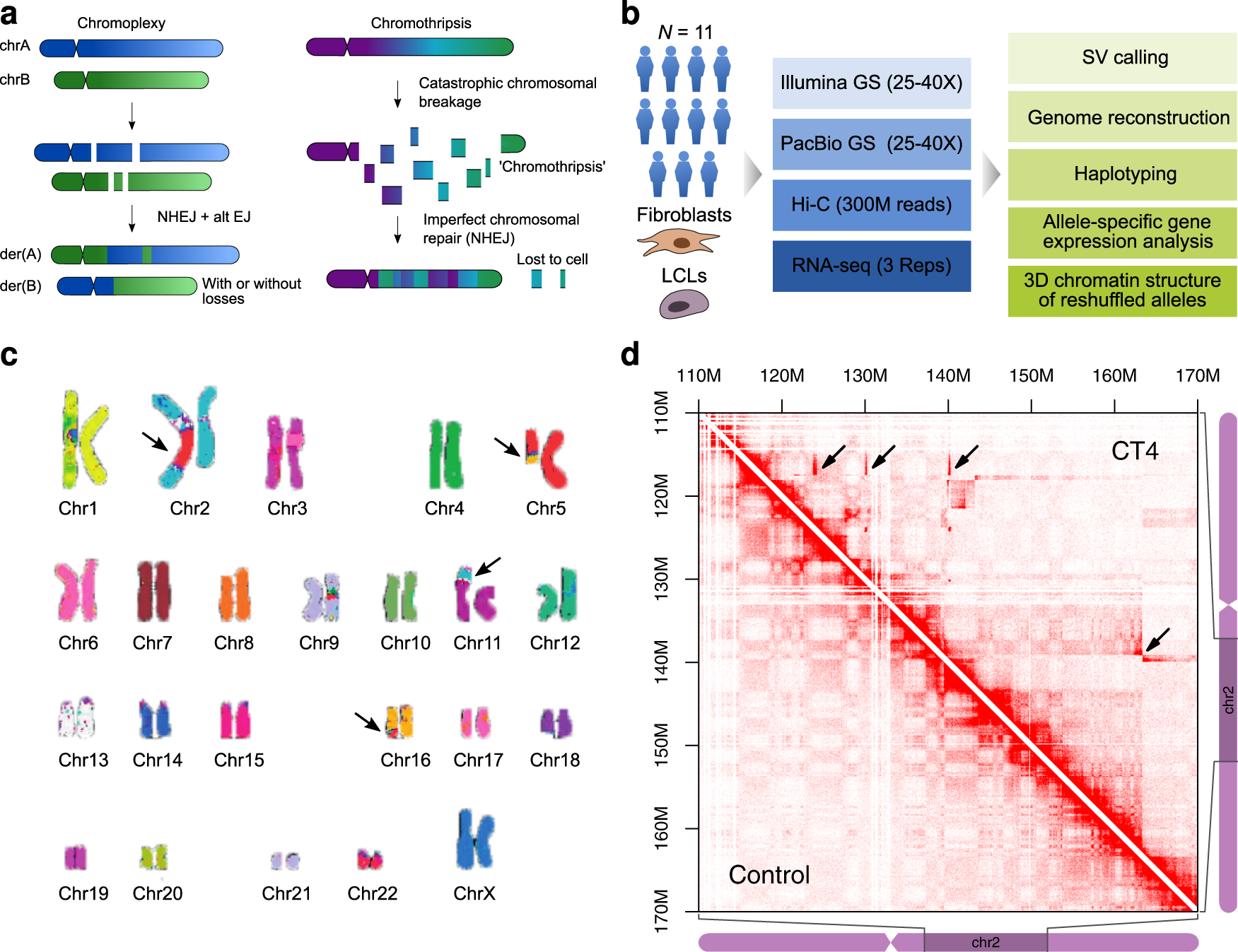
Integration of Hi-C with short and long-read genome sequencing reveals the structure of germline rearranged genomes | Nature Communications

Hi-C 2.0: An Optimized Hi-C Procedure for High-Resolution Genome-Wide Mapping of Chromosome Conformation | bioRxiv