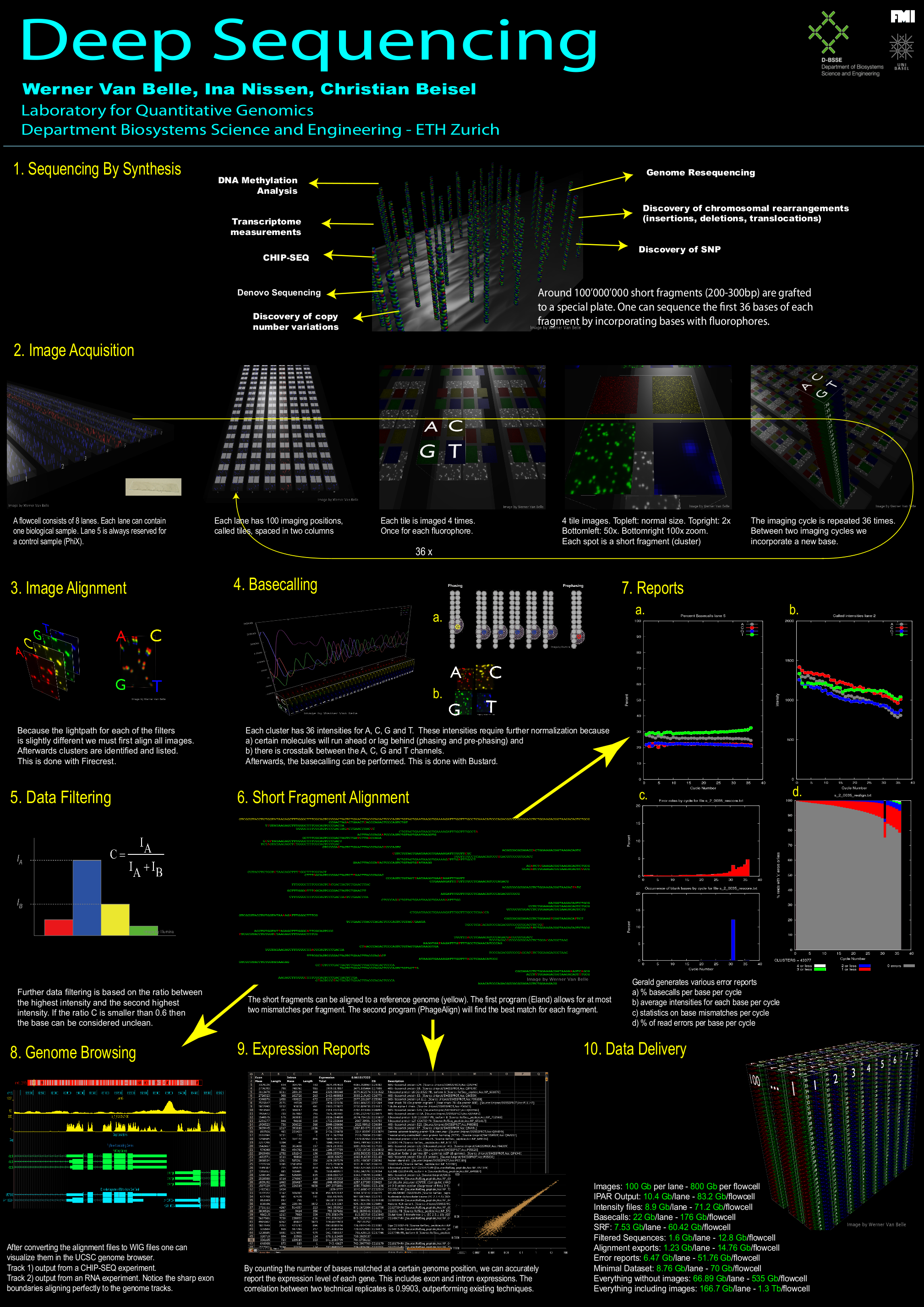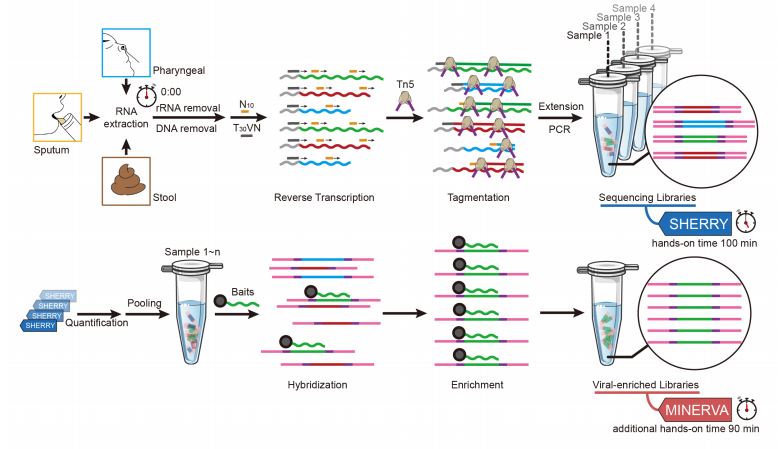
MINERVA: A facile strategy for SARS-CoV-2 whole genome deep sequencing of clinical samples - preLights

Metagenomic Next Generation Sequencing: How Does It Work and Is It Coming to Your Clinical Microbiology Lab?
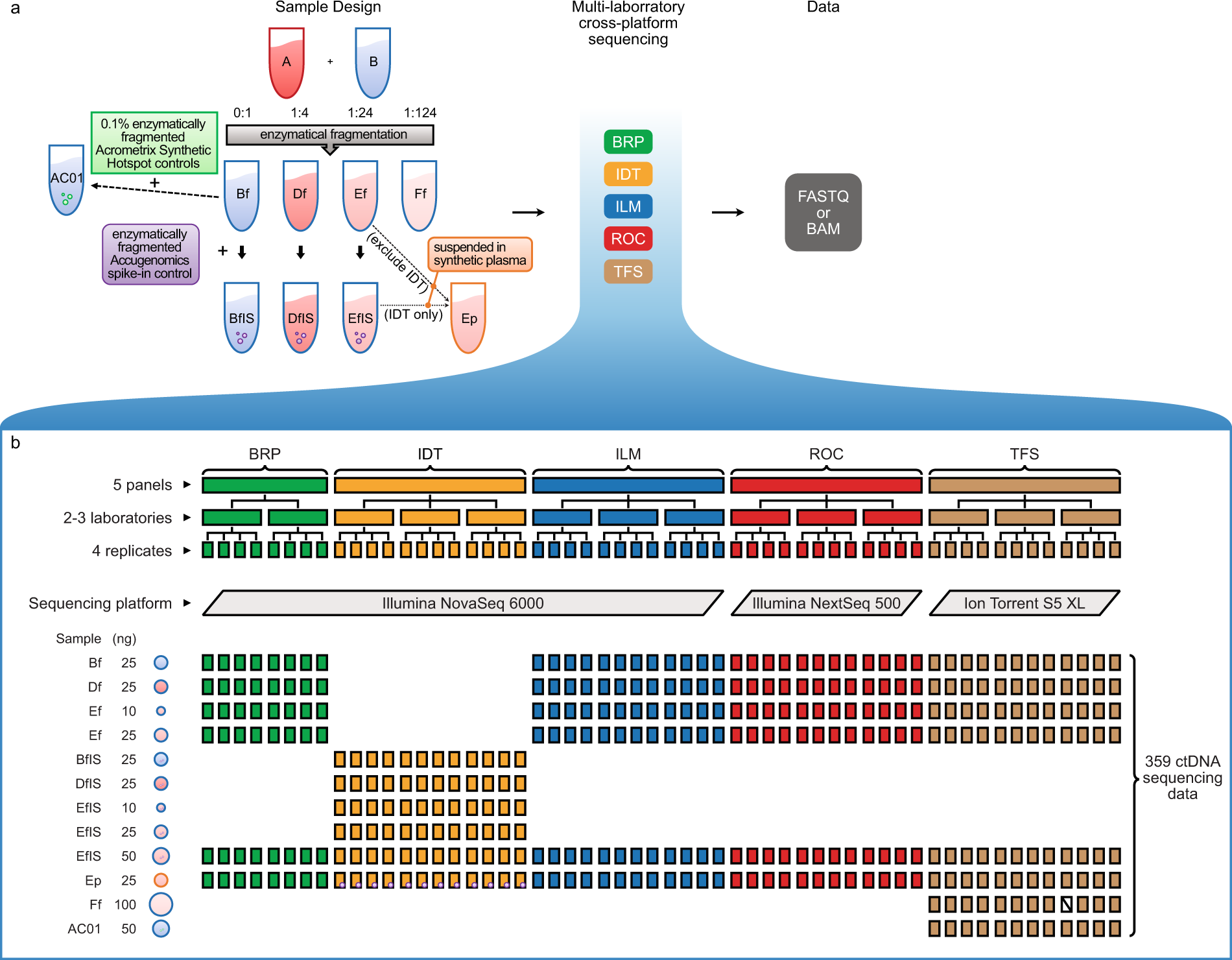
Ultra-deep sequencing data from a liquid biopsy proficiency study demonstrating analytic validity | Scientific Data
Strategy for deep sequencing full-length PSTVd progeny genomes. (A) RNA... | Download Scientific Diagram

Using deep sequencing to characterize the biophysical mechanism of a transcriptional regulatory sequence | PNAS
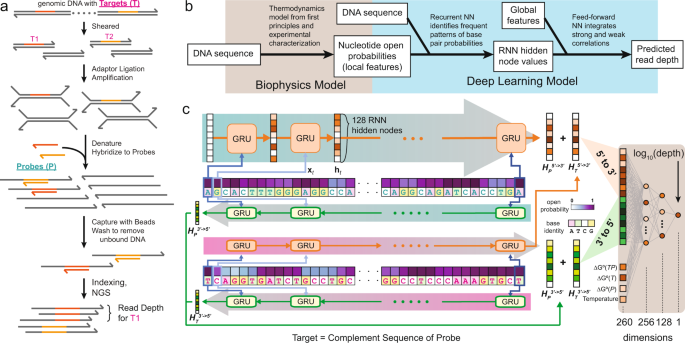
A deep learning model for predicting next-generation sequencing depth from DNA sequence | Nature Communications






![PDF] Making sense of deep sequencing. | Semantic Scholar PDF] Making sense of deep sequencing. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/cba4443cb5675cea52505a895538e6308aa9f0bf/3-Figure1-1.png)