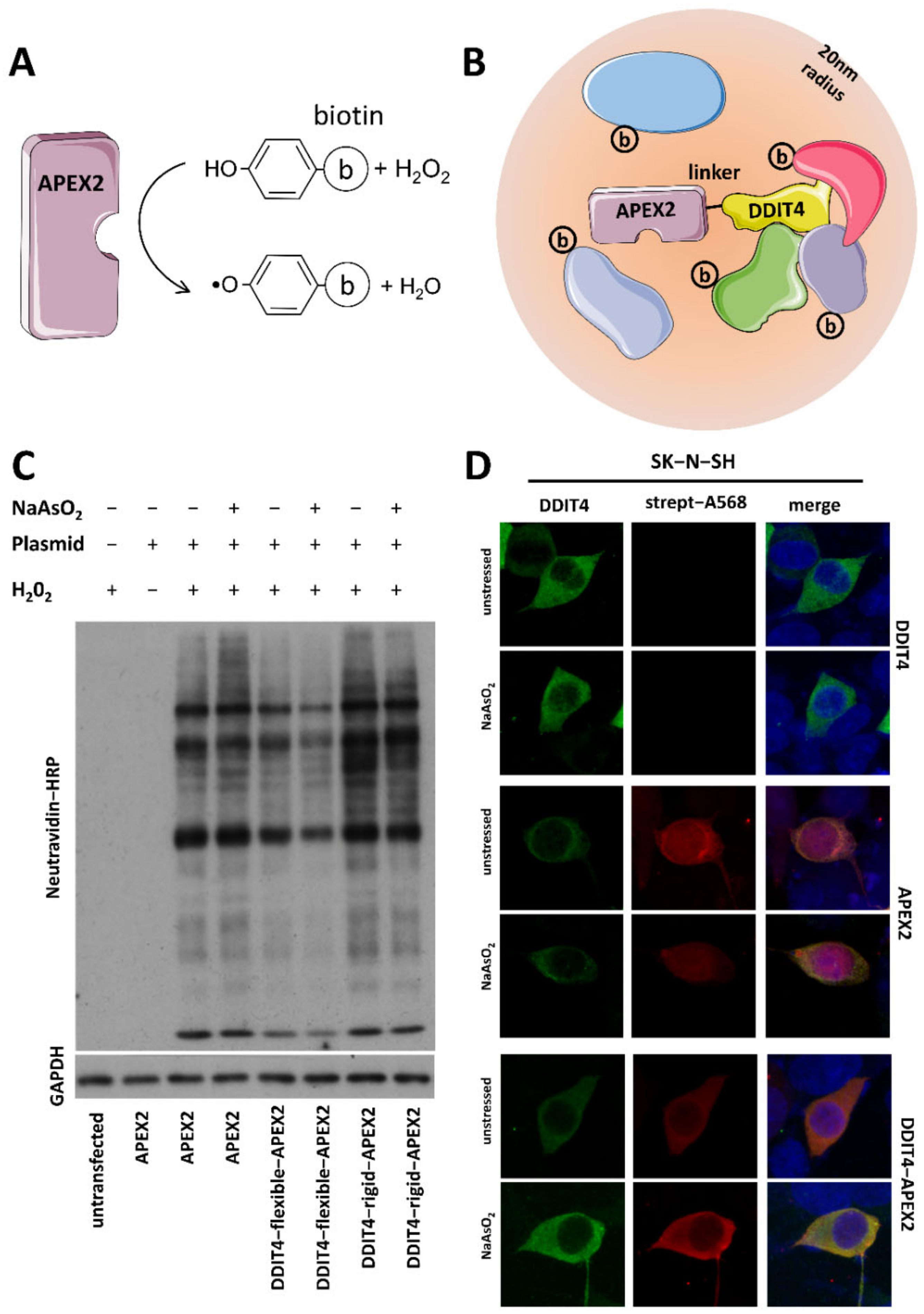
Proximity RNA Labeling by APEX-Seq Reveals the Organization of Translation Initiation Complexes and Repressive RNA Granules - ScienceDirect

APEX2‐mediated RAB proximity labeling identifies a role for RAB21 in clathrin‐independent cargo sorting | EMBO reports

Proximity RNA Labeling by APEX-Seq Reveals the Organization of Translation Initiation Complexes and Repressive RNA Granules - ScienceDirect
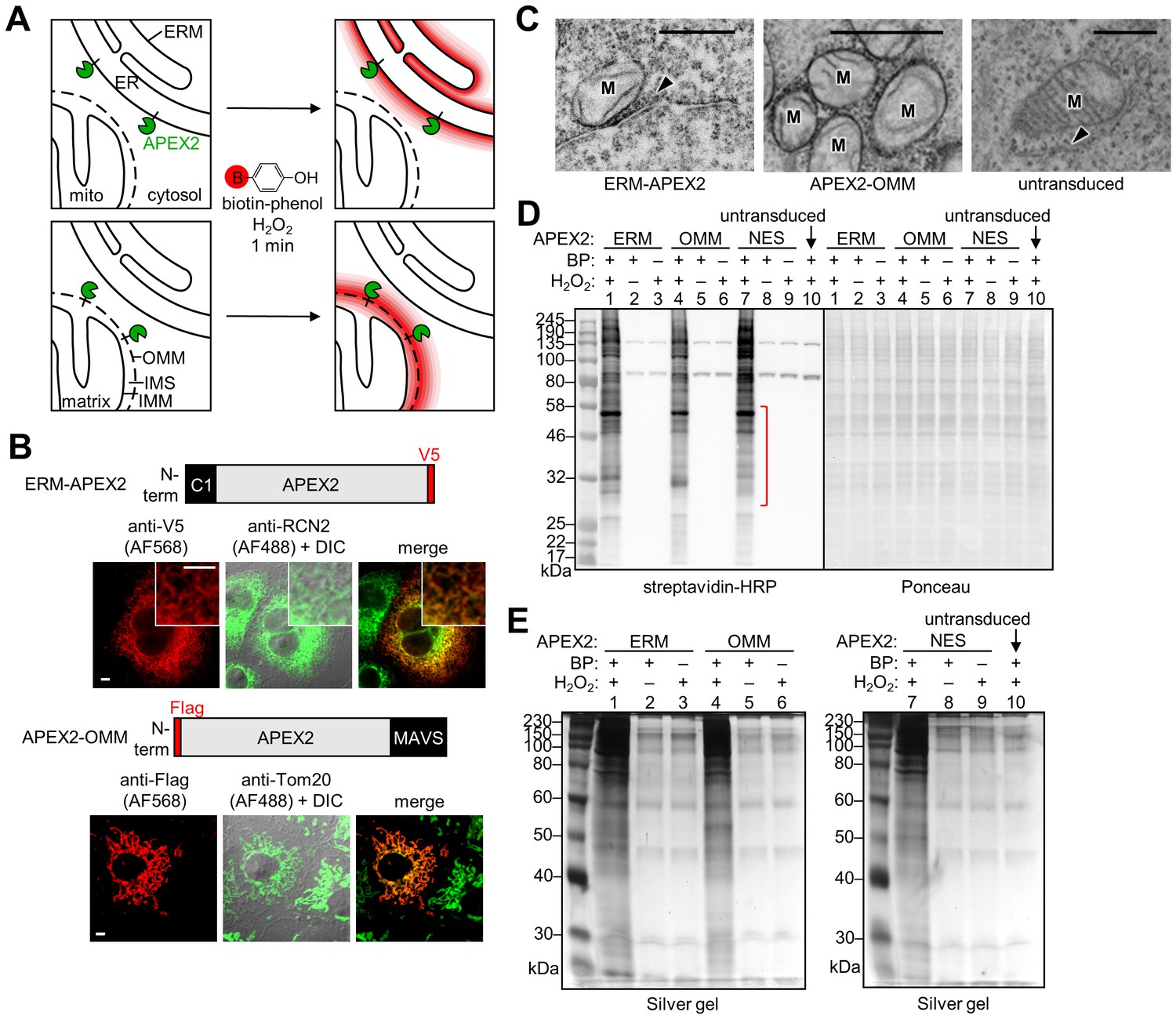
Proteomic mapping of cytosol-facing outer mitochondrial and ER membranes in living human cells by proximity biotinylation | eLife

Proximity-based labeling reveals DNA damage-induced N-terminal phosphorylation of fused in sarcoma (FUS) leads to distinct changes in the FUS protein interactome | bioRxiv
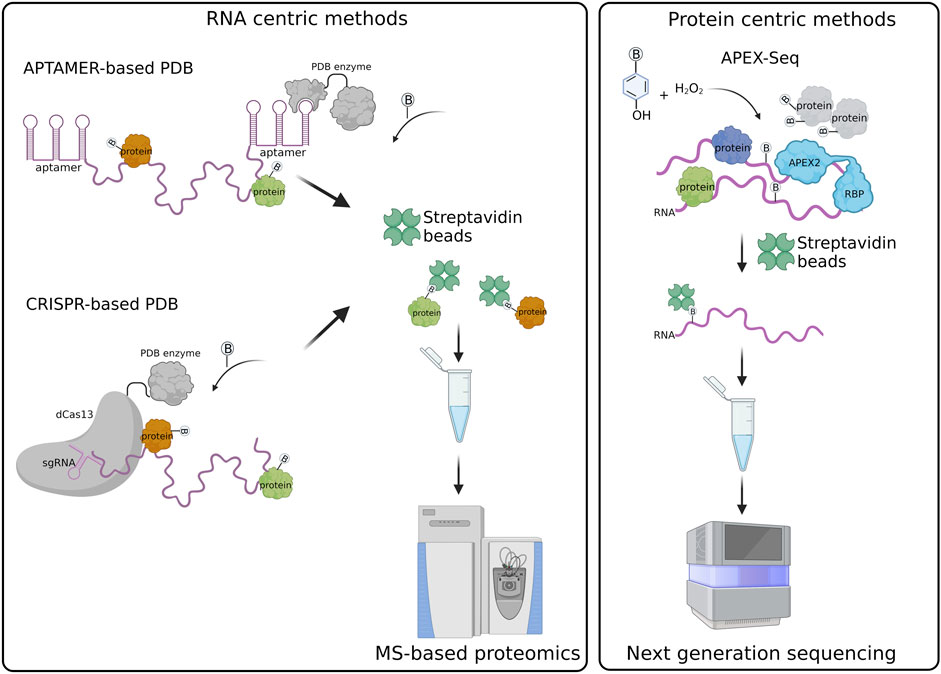
Frontiers | Proximity-dependent biotinylation technologies for mapping RNA-protein interactions in live cells

APEX2‐based Proximity Labeling of Atox1 Identifies CRIP2 as a Nuclear Copper‐binding Protein that Regulates Autophagy Activation - Chen - 2021 - Angewandte Chemie International Edition - Wiley Online Library

In vivo discovery of RNA proximal proteins in human cells via proximity-dependent biotinylation | bioRxiv
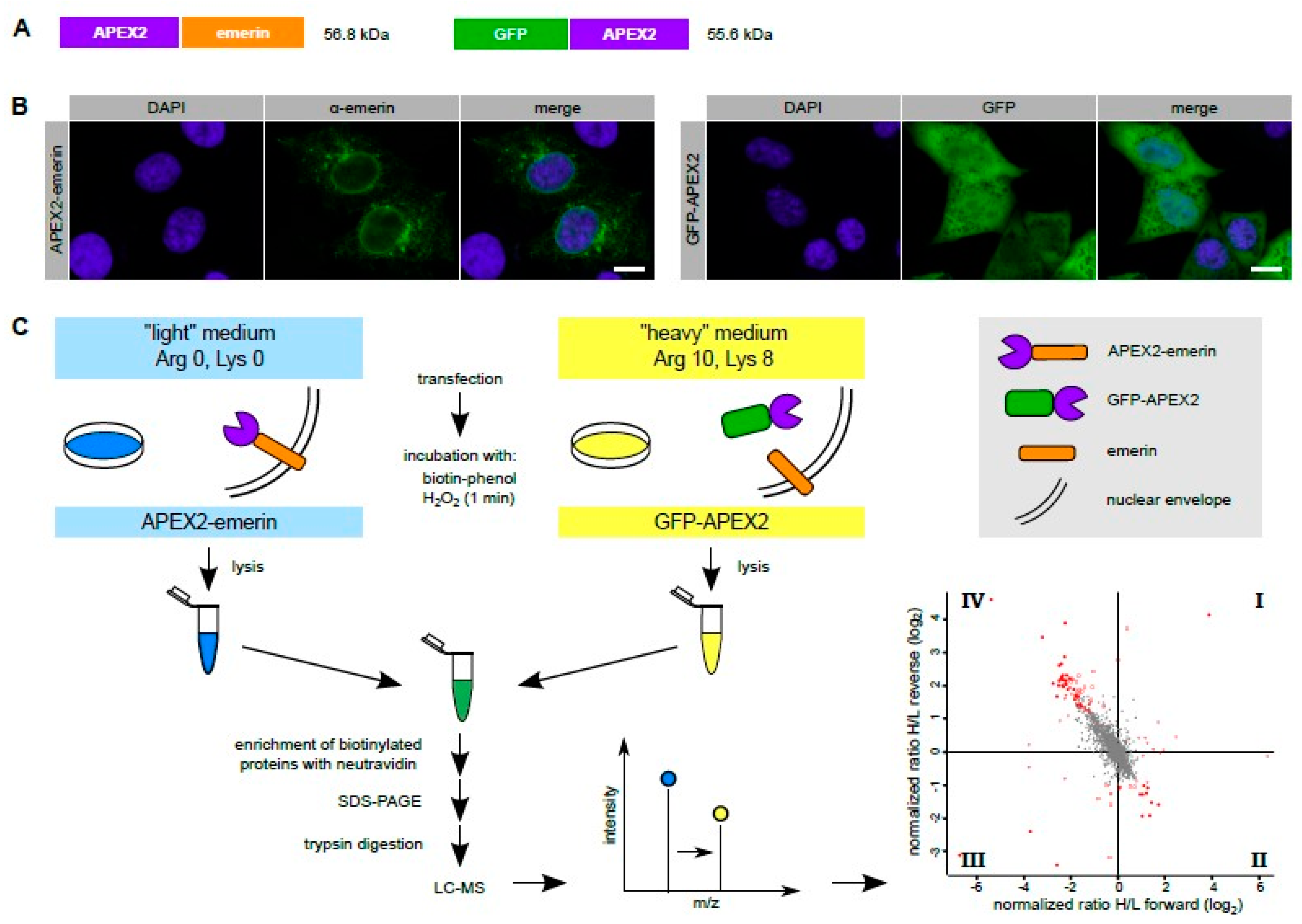
Cells | Free Full-Text | Probing the Environment of Emerin by Enhanced Ascorbate Peroxidase 2 (APEX2)-Mediated Proximity Labeling