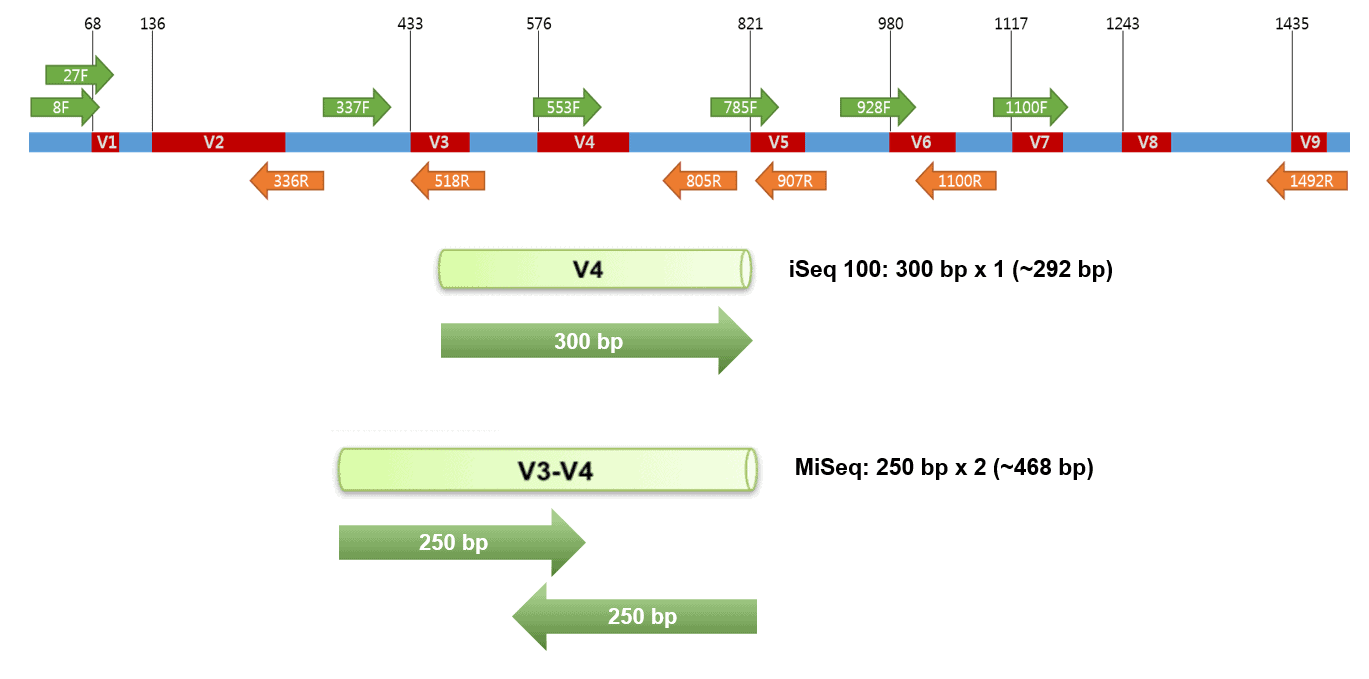
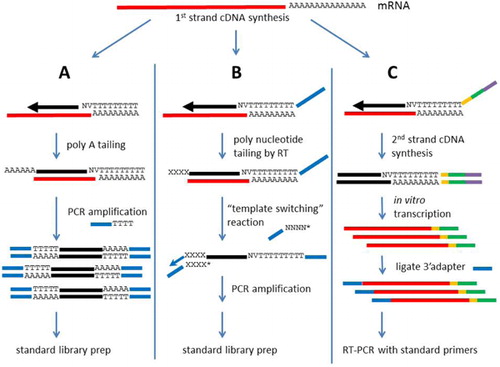
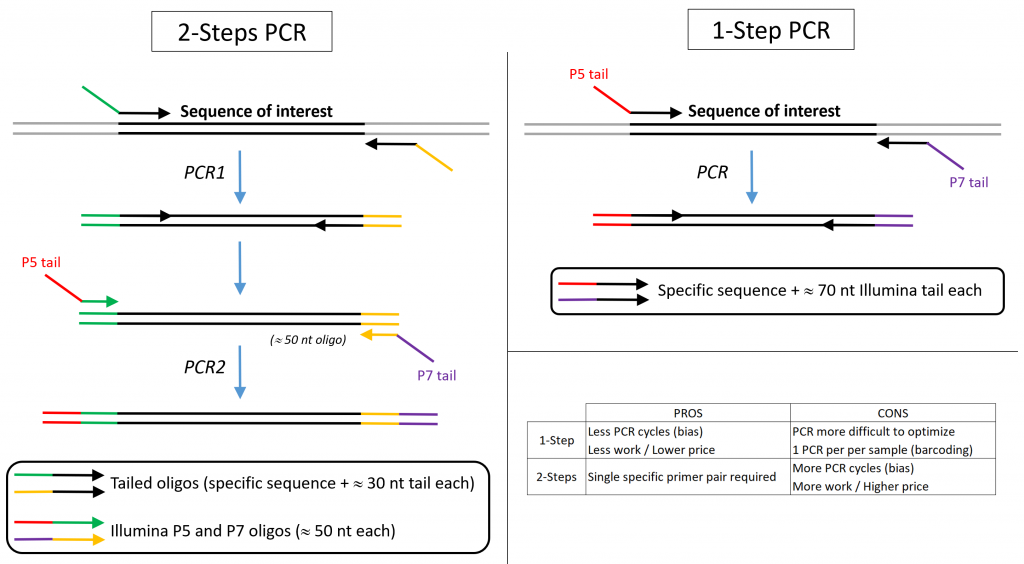
Targeted Amplicon Sequencing for Single-Nucleotide-Polymorphism Genotyping of Attaching and Effacing Escherichia coli O26:H11 Cattle Strains via a High-Throughput Library Preparation Technique | Applied and Environmental Microbiology

Different Amplicon Targets for Sequencing-Based Studies of Fungal Diversity | Applied and Environmental Microbiology
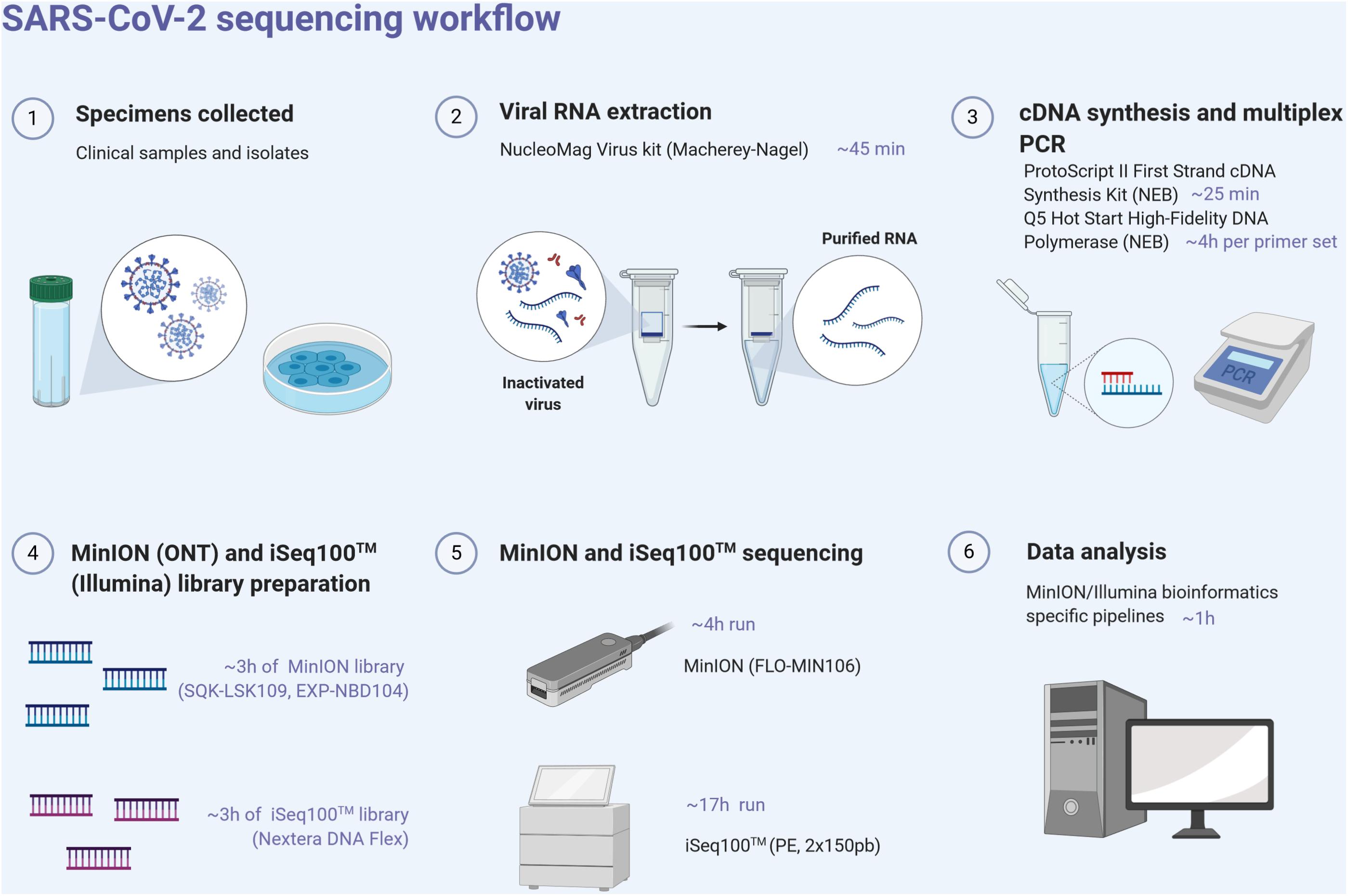
Frontiers | Rapid Genomic Characterization of SARS-CoV-2 by Direct Amplicon-Based Sequencing Through Comparison of MinION and Illumina iSeq100TM System
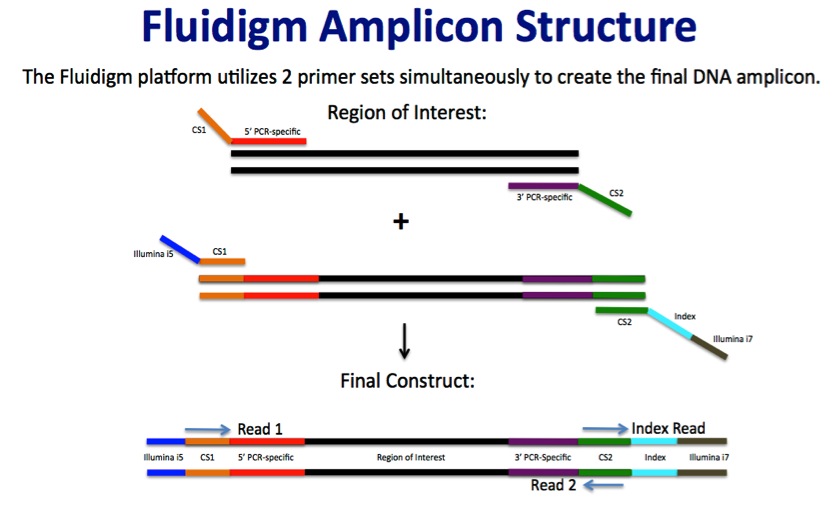
HTDNA: Services/Equipment: Preparation of 16S rRNA (and all other) Amplicons for Illumina Sequencing | Biotech
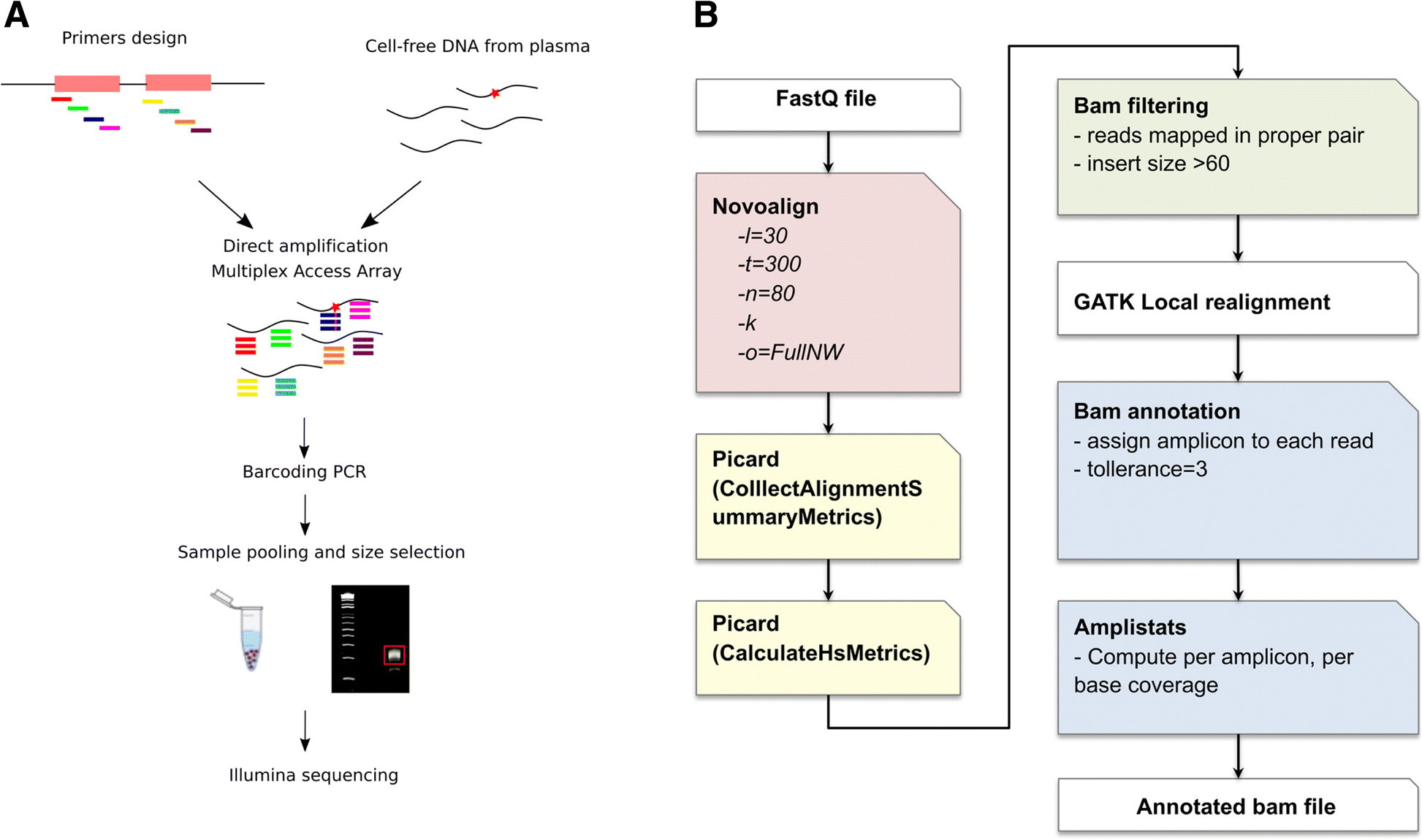
Next Generation-Targeted Amplicon Sequencing (NG-TAS): an optimised protocol and computational pipeline for cost-effective profiling of circulating tumour DNA | Genome Medicine | Full Text

An optimized, amplicon-based approach for sequencing of SARS-CoV-2 from patient samples using COVIDSeq assay on Illumina MiSeq sequencing platforms - ScienceDirect
Highly multiplexed gene amplicon sequencing | Joint Microbiome Facility of the Medical University of Vienna and the University of Vienna
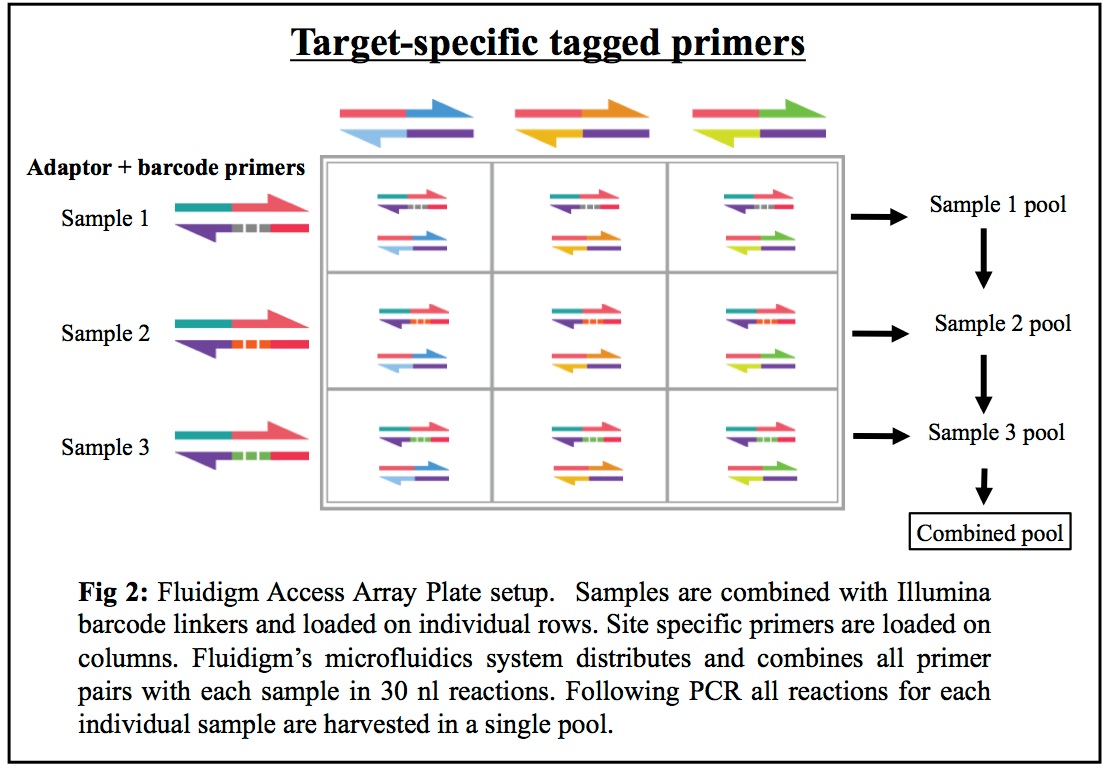
HTDNA: Services/Equipment: Preparation of 16S rRNA (and all other) Amplicons for Illumina Sequencing | Biotech
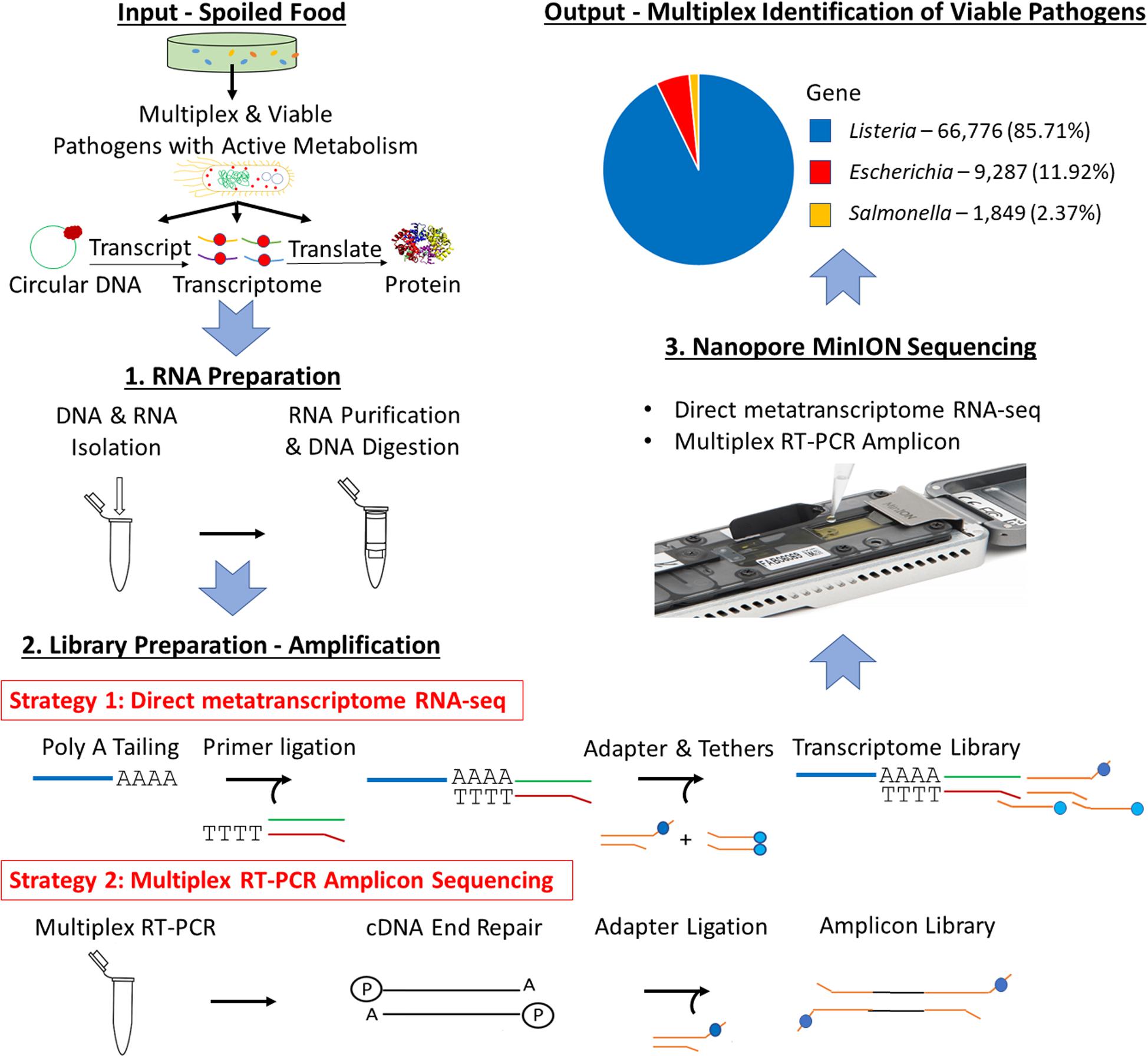
Frontiers | Direct Metatranscriptome RNA-seq and Multiplex RT-PCR Amplicon Sequencing on Nanopore MinION – Promising Strategies for Multiplex Identification of Viable Pathogens in Food
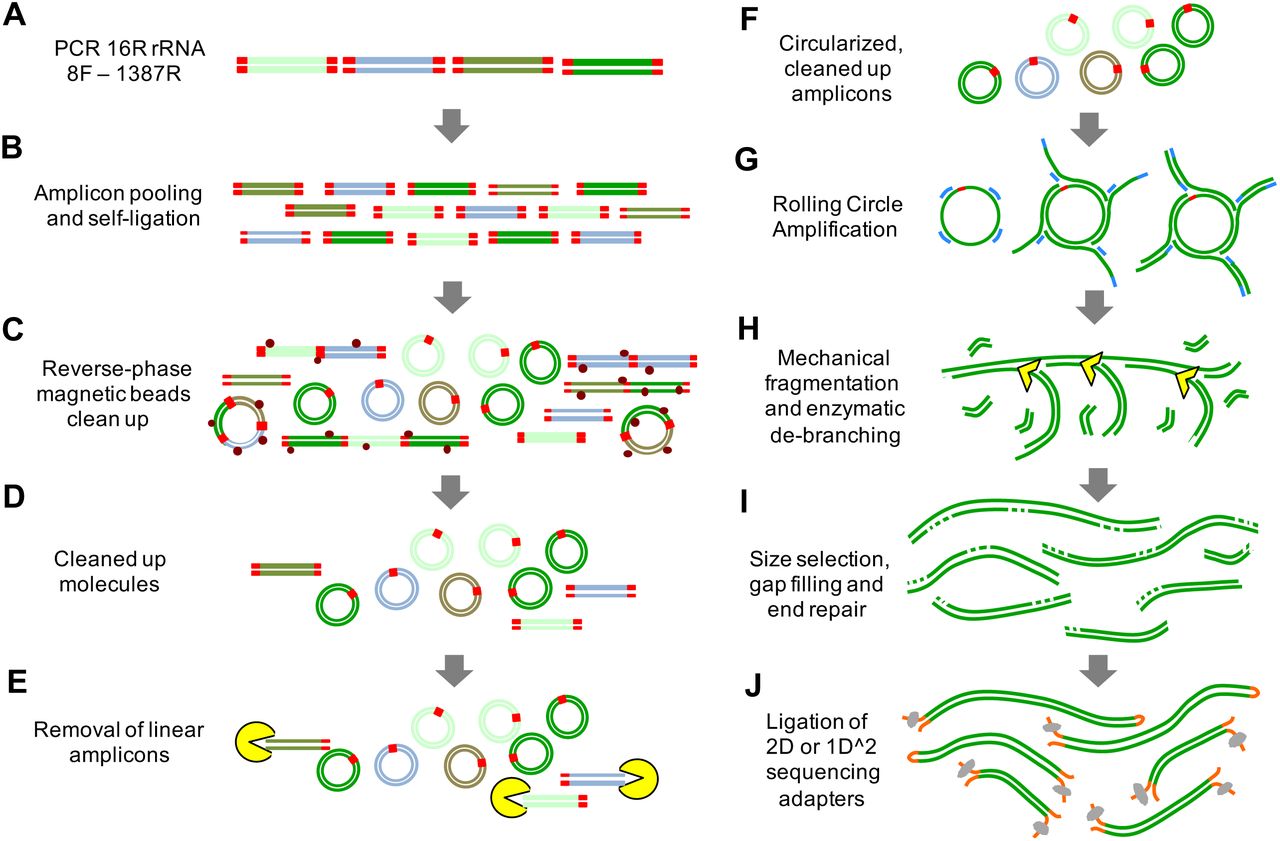
NanoAmpli-Seq: A workflow for amplicon sequencing from mixed microbial communities on the nanopore sequencing platform | bioRxiv

Population admixtures in medaka inferred by multiple arbitrary amplicon sequencing | Scientific Reports
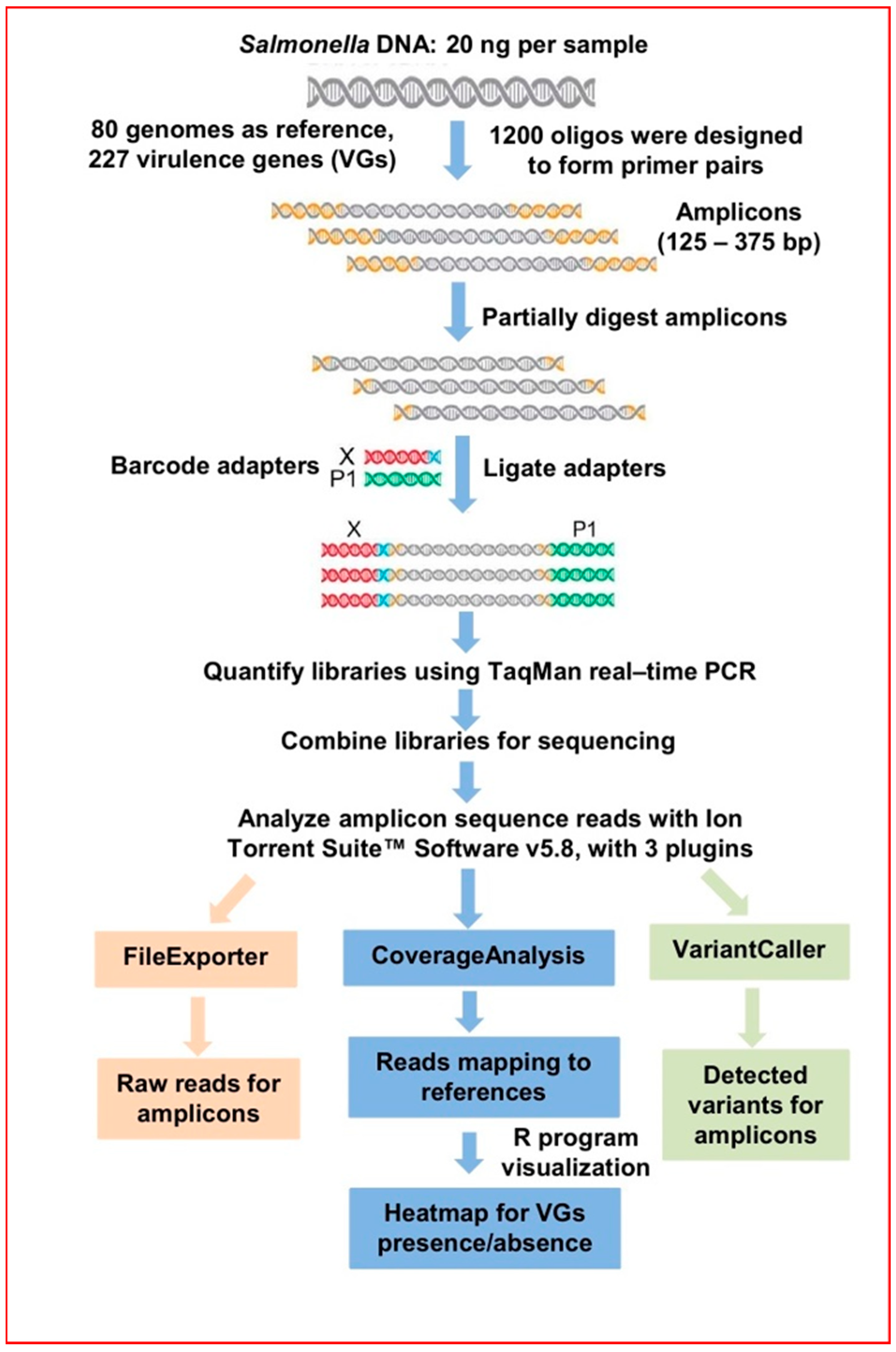
Microorganisms | Free Full-Text | Application of a High-Throughput Targeted Sequence AmpliSeq Procedure to Assess the Presence and Variants of Virulence Genes in Salmonella

![Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ] Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/7786/1/fig-1-full.png)