Figure 2 from and Richard Protease Poly ( A )-Binding Protein by Enterovirus 3 C Efficient Cleavage of Ribosome-Associated | Semantic Scholar

Human Rhinovirus 3C protease cleaves RIPK1, concurrent with caspase 8 activation | Scientific Reports

Whole-Genome Sequence Analysis Reveals the Enterovirus D68 Isolates during the United States 2014 Outbreak Mainly Belong to a Novel Clade | Scientific Reports
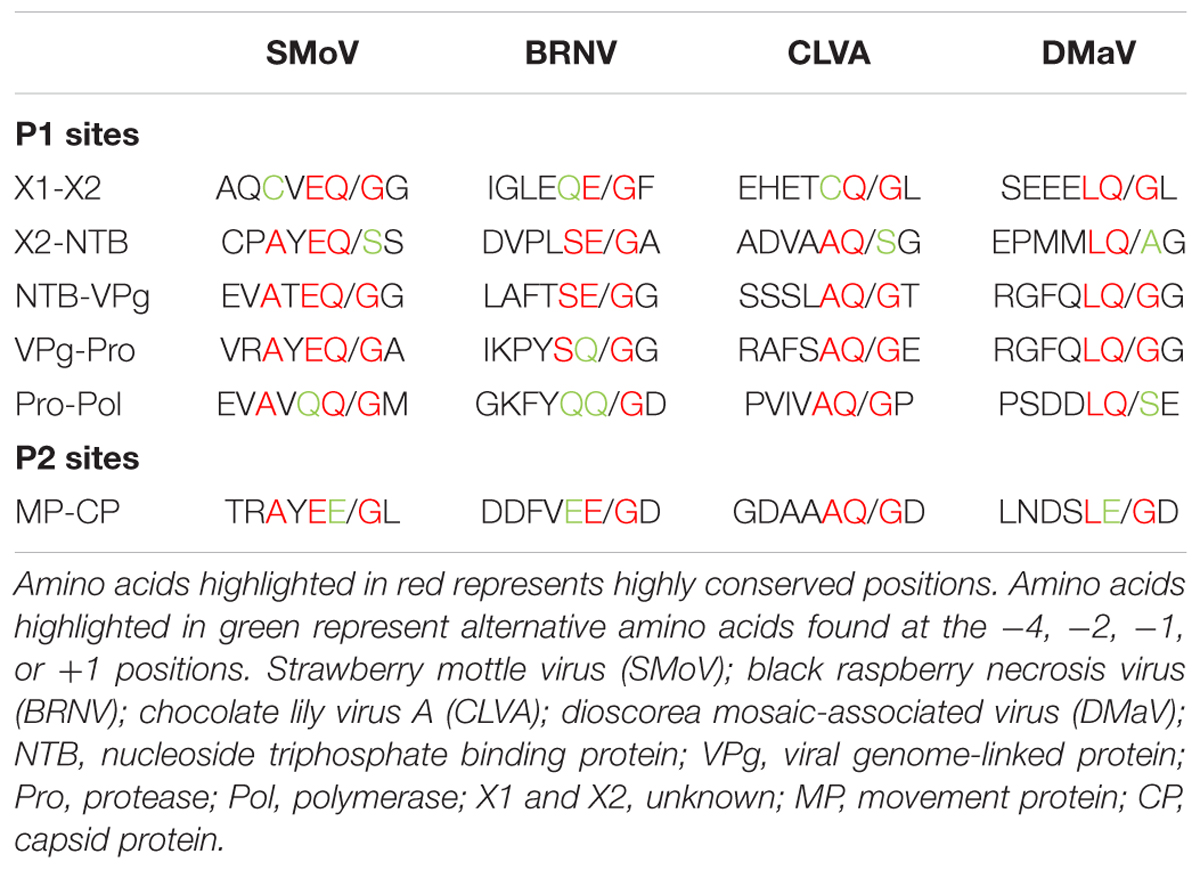

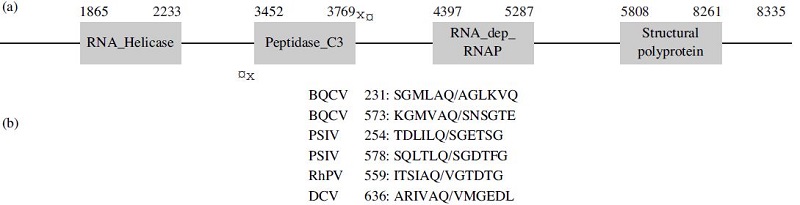
Frontiers | Identification of Cleavage Sites Recognized by the 3C-Like Cysteine Protease within the Two Polyproteins of Strawberry Mottle Virus
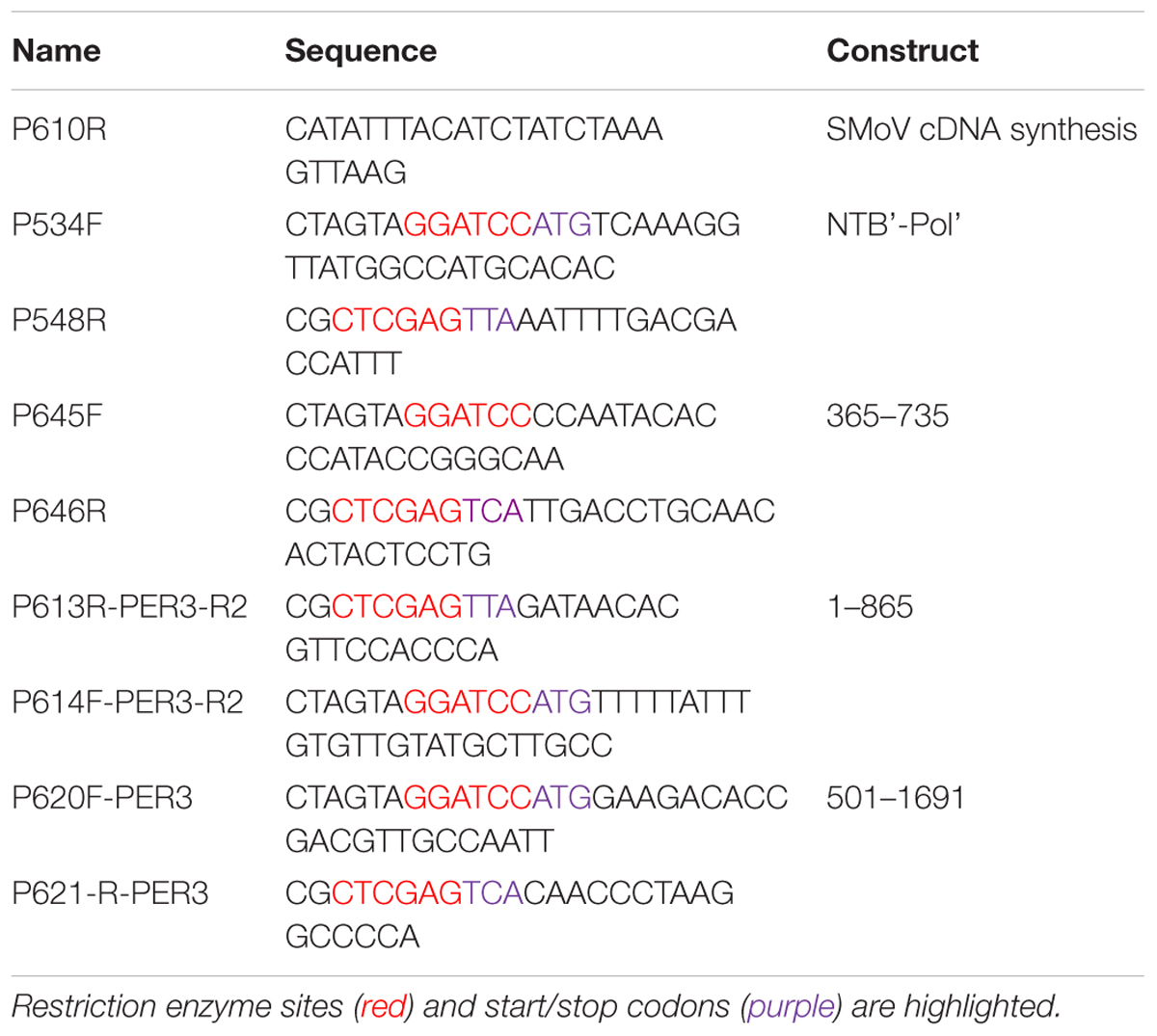
Frontiers | Identification of Cleavage Sites Recognized by the 3C-Like Cysteine Protease within the Two Polyproteins of Strawberry Mottle Virus

NT*-HRV3CP: An optimized construct of human rhinovirus 14 3C protease for high-yield expression and fast affinity-tag cleavage - ScienceDirect
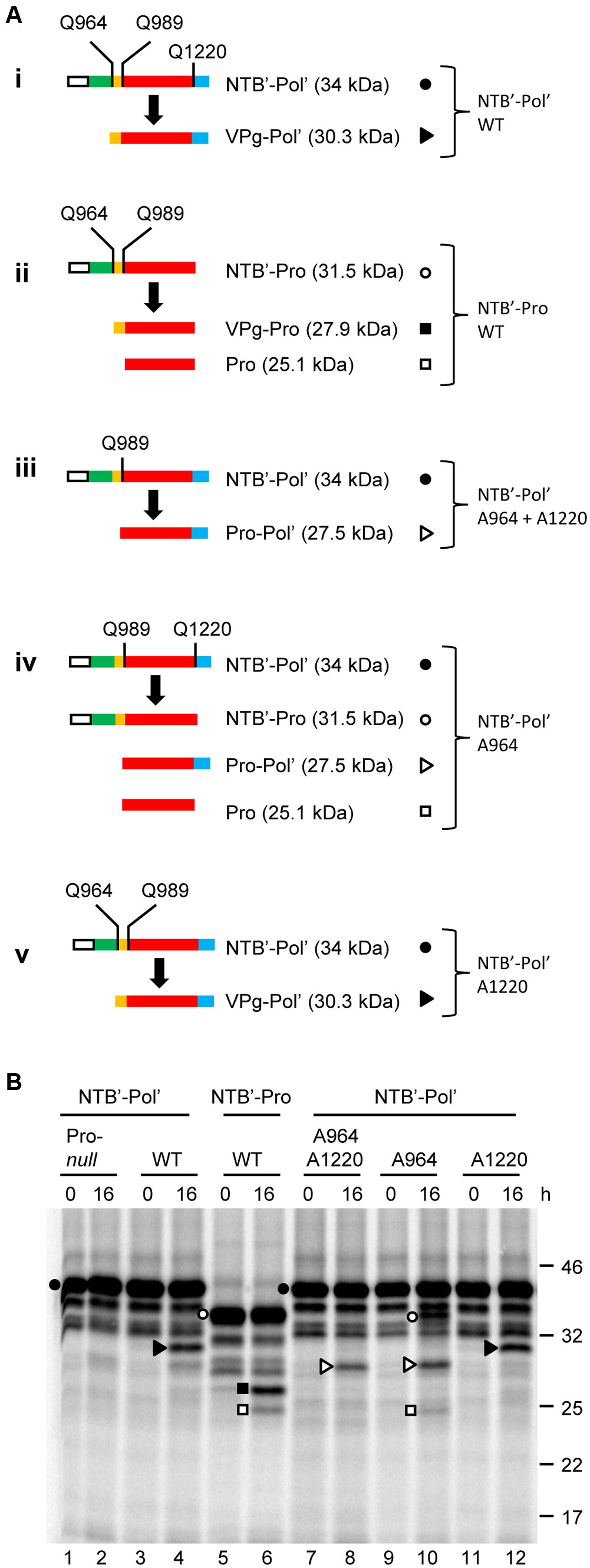
Frontiers | Identification of Cleavage Sites Recognized by the 3C-Like Cysteine Protease within the Two Polyproteins of Strawberry Mottle Virus
Determinants of the VP1/2A junction cleavage by the 3C protease in foot-and-mouth disease virus-infected cells | Microbiology Society

TMPRSS2 and furin are both essential for proteolytic activation of SARS-CoV-2 in human airway cells | Life Science Alliance

Quantitative Analysis of the Substrate Specificity of Human Rhinovirus 3C Protease and Exploration of Its Substrate Recognition Mechanisms | ACS Chemical Biology

Quantitative Analysis of the Substrate Specificity of Human Rhinovirus 3C Protease and Exploration of Its Substrate Recognition Mechanisms | ACS Chemical Biology
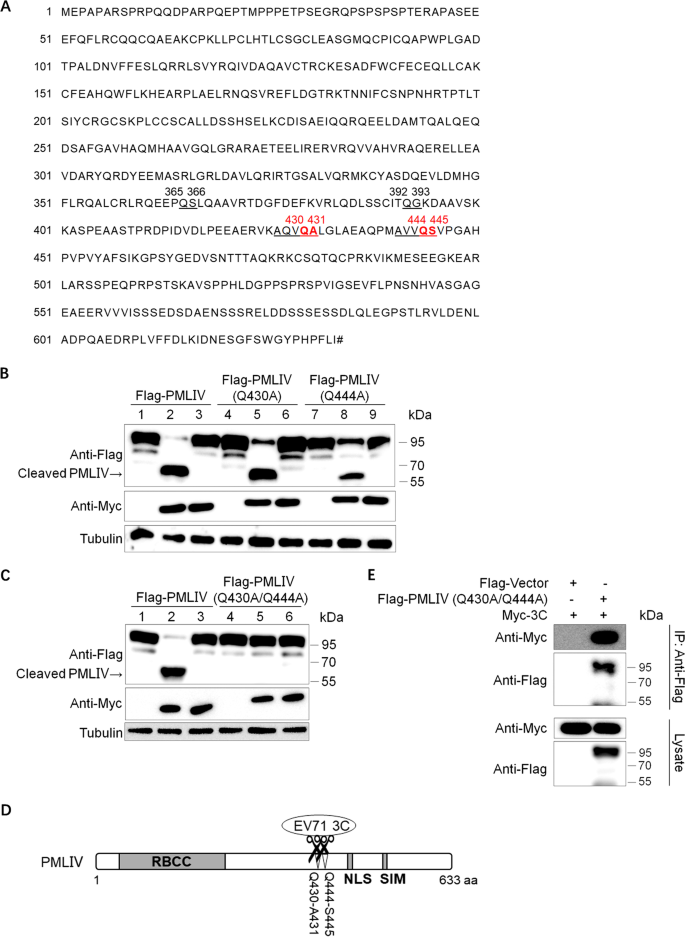
Enterovirus 71 3C Protease Cleaves a Novel Target CstF-64 and Inhibits Cellular Polyadenylation | PLOS Pathogens